Novel Aminoglycoside-Tolerant Phoenix Colony Variants of Pseudomonas aeruginosa
- PMID: 32540981
- PMCID: PMC7449220
- DOI: 10.1128/AAC.00623-20
Novel Aminoglycoside-Tolerant Phoenix Colony Variants of Pseudomonas aeruginosa
Abstract
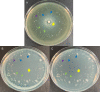
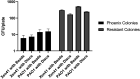
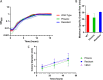
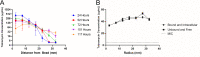
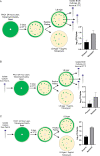
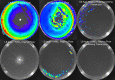
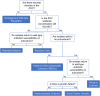
Pseudomonas aeruginosa is an opportunistic bacterial pathogen and is known to produce biofilms. We previously showed the emergence of colony variants in the presence of tobramycin-loaded calcium sulfate beads. In this study, we characterized the variant colonies, which survived the antibiotic treatment, and identified three distinct phenotypes-classically resistant colonies, viable but nonculturable colonies (VBNC), and phoenix colonies. Phoenix colonies, described here for the first time, grow out of the zone of clearance of antibiotic-loaded beads from lawn biofilms while there are still very high concentrations of antibiotic present, suggesting an antibiotic-resistant phenotype. However, upon subculturing of these isolates, phoenix colonies return to wild-type levels of antibiotic susceptibility. Compared with the wild type, phoenix colonies are morphologically similar aside from a deficiency in green pigmentation. Phoenix colonies do not recapitulate the phenotype of any previously described mechanisms of resistance, tolerance, or persistence and, thus, form a novel group with their own phenotype. Growth under anaerobic conditions suggests that an alternative metabolism could lead to the formation of phoenix colonies. These findings suggest that phoenix colonies could emerge in response to antibiotic therapies and lead to recurrent or persistent infections, particularly within biofilms where microaerobic or anaerobic environments are present.
Keywords: Pseudomonas aeruginosa; aminoglycosides; antibiotic tolerance; biofilms.
Copyright © 2020 American Society for Microbiology.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

