The differential strain virulence of the candidate toxins of Photorhabdus akhurstii can be correlated with their inter-strain gene sequence diversity
- PMID: 32550116
- PMCID: PMC7286996
- DOI: 10.1007/s13205-020-02288-0
The differential strain virulence of the candidate toxins of Photorhabdus akhurstii can be correlated with their inter-strain gene sequence diversity
Abstract
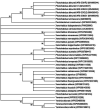
Photorhabdus akhurstii is an insect-parasitic bacterium that symbiotically associates with the nematode, Heterorhabditis indica. The bacterium possesses several pathogenicity islands that aids in conferring toxicity to different insects. Herein, we constructed the plasmid clones of coding sequences of four toxin genes (pirA, tcaA, tccA and tccC; each was isolated from four P. akhurstii strains IARI-SGMG3, IARI-SGGJ2, IARI-SGHR2 and IARI-SGMS1) in Escherichia coli and subsequently, their biological activity were investigated against the fourth-instar larvae of the model insect, Galleria mellonella via intra-hemocoel injection. Bioinformatics analyses indicated inter-strain amino acid sequence difference at several positions of the candidate toxins. In corroboration, differential insecticidal activity of the identical toxin protein (PirA, TcaA, TccA and TccC conferred 15-59, 27-100, 25-100 and 33-98% insect mortality, respectively, across the strains) derived from the different bacterial strains was observed, suggesting that the diverse gene pool in Indian strains of P. akhurstii leads to strain-specific virulence in this bacterium. These toxin candidates appear to be an attractive option to deploy them in biopesticide development for managing the insect pests globally.
Keywords: Bacterial strains; Galleria mellonella; PirA; RVA assay; TcaA; TccA; TccC.
© King Abdulaziz City for Science and Technology 2020.
Figures









Similar articles
-
TcaB, an insecticidal protein from Photorhabdus akhurstii causes cytotoxicity in the greater wax moth, Galleria mellonella.Pestic Biochem Physiol. 2019 Jun;157:219-229. doi: 10.1016/j.pestbp.2019.03.019. Epub 2019 Apr 1. Pestic Biochem Physiol. 2019. PMID: 31153472
-
Identification of Galtox, a new protein toxin from Photorhabdus bacterial symbionts of Heterorhabditis nematodes.Toxicon. 2021 Apr 30;194:53-62. doi: 10.1016/j.toxicon.2021.02.011. Epub 2021 Feb 18. Toxicon. 2021. PMID: 33610634
-
A toxin complex protein from Photorhabdus akhurstii conferred oral insecticidal activity against Galleria mellonella by targeting the midgut epithelium.Microbiol Res. 2021 Jan;242:126642. doi: 10.1016/j.micres.2020.126642. Epub 2020 Nov 5. Microbiol Res. 2021. PMID: 33191102
-
A bacterial binary toxin system that kills both insects and aquatic crustaceans: Photorhabdus insect-related toxins A and B.PLoS Pathog. 2023 May 4;19(5):e1011330. doi: 10.1371/journal.ppat.1011330. eCollection 2023 May. PLoS Pathog. 2023. PMID: 37141203 Free PMC article. Review.
-
Insecticidal toxins from Photorhabdus bacteria and their potential use in agriculture.Toxicon. 2007 Mar 15;49(4):436-51. doi: 10.1016/j.toxicon.2006.11.019. Epub 2006 Nov 30. Toxicon. 2007. PMID: 17207509 Review.
Cited by
-
Photorhabdus spp.: An Overview of the Beneficial Aspects of Mutualistic Bacteria of Insecticidal Nematodes.Plants (Basel). 2021 Aug 12;10(8):1660. doi: 10.3390/plants10081660. Plants (Basel). 2021. PMID: 34451705 Free PMC article. Review.
-
The induced knockdown of GmCAD receptor protein encoding gene in Galleria mellonella decreased the insect susceptibility to a Photorhabdus akhurstii oral toxin.Virulence. 2021 Dec;12(1):2957-2971. doi: 10.1080/21505594.2021.2006996. Virulence. 2021. PMID: 34882066 Free PMC article.
References
-
- Aloy P, Russell RB. Interprets: protein interaction prediction through tertiary structure. Bioinformatics. 2003;19:161–162. - PubMed
-
- Boemare N. Interactions between the partners of the entomopathogenic bacterium nematode complexes, Steinernema-Xenorhabdus and Heterorhabditis-Photorhabdus. Nematology. 2002;4:601–603.
-
- Bowen D, Rocheleau TA, Blackburn M, Andreev O, Golubeva E, Bhartiaffrench-Constant RRH. Insecticidal toxins from the bacterium Photorhabdus luminescens. Science. 1998;280:2129–2132. - PubMed
LinkOut - more resources
Full Text Sources

