Protection against COVID-19 injury by qingfei paidu decoction via anti-viral, anti-inflammatory activity and metabolic programming
- PMID: 32554251
- PMCID: PMC7247521
- DOI: 10.1016/j.biopha.2020.110281
Protection against COVID-19 injury by qingfei paidu decoction via anti-viral, anti-inflammatory activity and metabolic programming
Abstract
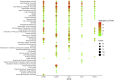
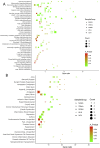
Qingfei Paidu decoction (QFPD), a multi-component herbal formula, has been widely used to treat COVID-19 in China. However, its active compounds and mechanisms of action are still unknown. Firstly, we divided QFPD into five functional units (FUs) according to the compatibility theory of traditional Chinese medicine. The corresponding common targets of the five FUs were all significantly enriched in Go Ontology (oxidoreductase activity, lipid metabolic process, homeostatic process, etc.), KEGG pathways (steroid biosynthesis, PPAR signaling pathway, adipocytokine signaling pathway, etc.), TTD diseases (chronic inflammatory diseases, asthma, chronic obstructive pulmonary Disease, etc.), miRNA (MIR183), kinase (CDK7) and TF (LXR). QFPD contained 257 specific targets in addition to HCoV, pneumonia and ACE2 co-expression proteins. Then, network topology analysis of the five components-target-pathway-disease networks yielded 67 active ingredients. In addition, ADMET estimations showed that 20 compounds passed the stringent lead-like criteria and in silico drug-likeness test with high gastrointestinal absorption and the median lethal dose (LD50 > 1600 mg/kg). Moreover, 4 specific ingredients (M3, S1, X2 and O2) and 5 common ingredients (MS1, MX16, SX1, WO1 and XO1) of QFPD presented good molecular docking score for 2019-nCov structure and non-structure proteins. Finally, drug perturbation of COVID-19 network robustness showed that all five FUs may protect COVID-19 independently, and target 8 specifically expressed drug-attacked nodes which were related to the bacterial and viral responses, immune system, signaling transduction, etc. In conclusion, our new FUNP analysis showed that QFPD had a protection effect on COVID-19 by regulating a complex molecular network with safety and efficacy. Part of the mechanism was associated with the regulation of anti-viral, anti-inflammatory activity and metabolic programming.
Keywords: Anti-inflammatory; Anti-viral; COVID-19; Functional units of network pharmacology; Metabolic programming; Qingfei paidu decoction.
Copyright © 2020 The Authors. Published by Elsevier Masson SAS.. All rights reserved.
Conflict of interest statement
The authors declare no conflict of interest.
Figures











References
-
- Doctor D.D. 2020. COVID-19 Global Pandemic Real-time Report.http://ncov.dxy.cn/ncovh5/view/en_pneumonia?from=dxy&source=dxys
-
- Hsieh C.F., Lo C.W., Liu C.H., Lin S., Yen H.R., Lin T.Y., Horng J.T. Mechanism by which ma-xing-shi-gan-tang inhibits the entry of influenza virus. J. Ethnopharmacol. 2012;143(1):57–67. - PubMed
-
- Lin C.C., Wang Y.Y., Chen S.M., Liu Y.T., Li J.Q., Li F., Dai J.C., Zhang T., Qiu F., Liu H., Dai Z., Zhang Z.D. Shegan-Mahuang Decoction ameliorates asthmatic airway hyperresponsiveness by downregulating Th2/Th17 cells but upregulating CD4+FoxP3+ Tregs. J. Ethnopharmacol. 2020;253 - PubMed
-
- Cheng P.W., Ng L.T., Lin C.C. Xiao chai hu tang inhibits CVB1 virus infection of CCFS-1 cells through the induction of Type I interferon expression. Int. Immunopharmacol. 2006;6(6):1003–1012. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

