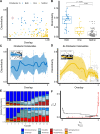
Dissimilarity-Overlap analysis of replicate enrichment communities
- PMID: 32555503
- PMCID: PMC7490387
- DOI: 10.1038/s41396-020-0702-7
Dissimilarity-Overlap analysis of replicate enrichment communities
Abstract
The taxonomic composition of microbial communities can vary substantially across habitats and within the same habitat over time. Efforts to build quantitative and predictive models of microbial population dynamics are underway, but fundamental questions remain. How different are population dynamics in different environments? Do communities that share the same taxa also exhibit identical dynamics? In vitro communities can help establish baseline expectations that are critical towards resolving these questions in natural communities. Here, we applied a recently developed tool, Dissimilarity-Overlap Analysis (DOA), to a set of experimental in vitro communities that differed in nutrient composition. The Dissimilarity and Overlap of these communities are negatively correlated in replicate habitats, as one would expect if microbial population dynamics were on average strongly convergent (or "universal") across these replicate habitats. However, the existence of such a negative correlation does not necessarily imply that population dynamics are always universal in all communities. Even in replicate, identical habitats, two different communities may contain the same set of taxa at different abundances in equilibrium. The formation of alternative states in community assembly is strongly associated with the presence of specific taxa in the communities. Our results benchmark DOA, providing support for some of its core assumptions, and suggest that communities sharing the same taxa and external abiotic factors generally (but not necessarily) have a negative correlation between Dissimilarity and Overlap.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures