Novel partiti-like viruses are conditional mutualistic symbionts in their normal lepidopteran host, African armyworm, but parasitic in a novel host, Fall armyworm
- PMID: 32569314
- PMCID: PMC7332103
- DOI: 10.1371/journal.ppat.1008467
Novel partiti-like viruses are conditional mutualistic symbionts in their normal lepidopteran host, African armyworm, but parasitic in a novel host, Fall armyworm
Abstract
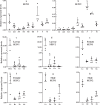
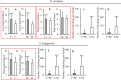
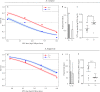
Recent advances in next generation sequencing (NGS) (e.g. metagenomic and transcriptomic sequencing) have facilitated the discovery of a large number of new insect viruses, but the characterization of these viruses is still in its infancy. Here, we report the discovery, using RNA-seq, of three new partiti-like viruses from African armyworm, Spodoptera exempta (Lepidoptera: Noctuidae), which are all vertically-transmitted transovarially from mother to offspring with high efficiency. Experimental studies show that the viruses reduce their host's growth rate and reproduction, but enhance their resistance to a nucleopolyhedrovirus (NPV). Via microinjection, these partiti-like viruses were transinfected into a novel host, a newly-invasive crop pest in sub-Saharan Africa (SSA), the Fall armyworm, S. frugiperda. This revealed that in this new host, these viruses appear to be deleterious without any detectable benefit; reducing their new host's reproductive rate and increasing their susceptibility to NPV. Thus, the partiti-like viruses appear to be conditional mutualistic symbionts in their normal host, S. exempta, but parasitic in the novel host, S. frugiperda. Transcriptome analysis of S. exempta and S. frugiperda infected, or not, with the partiti-like viruses indicates that the viruses may regulate pathways related to immunity and reproduction. These findings suggest a possible pest management strategy via the artificial host-shift of novel viruses discovered by NGS.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Partiti-like viruses from African armyworm increase larval and pupal mortality of a novel host: the Egyptian cotton leafworm.Pest Manag Sci. 2022 Apr;78(4):1529-1537. doi: 10.1002/ps.6771. Epub 2022 Jan 10. Pest Manag Sci. 2022. PMID: 34965003
-
The distribution of covert microbial natural enemies of a globally invasive crop pest, fall armyworm, in Africa: Enemy release and spillover events.J Anim Ecol. 2022 Sep;91(9):1826-1841. doi: 10.1111/1365-2656.13760. Epub 2022 Jun 22. J Anim Ecol. 2022. PMID: 35678697 Free PMC article.
-
High levels of genetic diversity in Spodoptera exempta NPV from Tanzania.J Invertebr Pathol. 2010 Oct;105(2):190-3. doi: 10.1016/j.jip.2010.06.008. Epub 2010 Jun 22. J Invertebr Pathol. 2010. PMID: 20600096
-
Viruses of the Fall Armyworm Spodoptera frugiperda: A Review with Prospects for Biological Control.Viruses. 2021 Nov 4;13(11):2220. doi: 10.3390/v13112220. Viruses. 2021. PMID: 34835026 Free PMC article. Review.
-
RNA interference (RNAi) applications to the management of fall armyworm, Spodoptera frugiperda (Lepidoptera: Noctuidae): Its current trends and future prospects.Front Mol Biosci. 2022 Sep 7;9:944774. doi: 10.3389/fmolb.2022.944774. eCollection 2022. Front Mol Biosci. 2022. PMID: 36158573 Free PMC article. Review.
Cited by
-
Insights into insecticide-resistance mechanisms in invasive species: Challenges and control strategies.Front Physiol. 2023 Jan 9;13:1112278. doi: 10.3389/fphys.2022.1112278. eCollection 2022. Front Physiol. 2023. PMID: 36699674 Free PMC article. Review.
-
Transcriptomic Investigation of the Virus Spectrum Carried by Midges in Border Areas of Yunnan Province.Viruses. 2024 Apr 25;16(5):674. doi: 10.3390/v16050674. Viruses. 2024. PMID: 38793556 Free PMC article.
-
Transcriptome Analysis Reveals a Diverse Range of Novel Viruses in Australian Sugarcane Soldier Fly (Inopus flavus) Larvae.Viruses. 2024 Mar 27;16(4):516. doi: 10.3390/v16040516. Viruses. 2024. PMID: 38675859 Free PMC article.
-
Infection pattern of male-killing viruses alters phenotypes in the tea tortrix moth Homona magnanima.Heredity (Edinb). 2025 Feb;134(2):120-128. doi: 10.1038/s41437-024-00741-x. Epub 2024 Dec 26. Heredity (Edinb). 2025. PMID: 39725691
-
Diversity and transmissibility of RNA viruses in the small brown planthopper, Laodelphax striatellus.J Virol. 2024 Dec 17;98(12):e0019124. doi: 10.1128/jvi.00191-24. Epub 2024 Nov 26. J Virol. 2024. PMID: 39589138 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

