Development of Cellular Models to Study Efficiency and Safety of Gene Edition by Homologous Directed Recombination Using the CRISPR/Cas9 System
- PMID: 32570971
- PMCID: PMC7349026
- DOI: 10.3390/cells9061492
Development of Cellular Models to Study Efficiency and Safety of Gene Edition by Homologous Directed Recombination Using the CRISPR/Cas9 System
Abstract
In spite of the enormous potential of CRISPR/Cas in basic and applied science, the levels of undesired genomic modifications cells still remain mostly unknown and controversial. Nowadays, the efficiency and specificity of the cuts generated by CRISPR/Cas is the main concern. However, there are also other potential drawbacks when DNA donors are used for gene repair or gene knock-ins. These GE strategies should take into account not only the specificity of the nucleases, but also the fidelity of the DNA donor to carry out their function. The current methods to quantify the fidelity of DNA donor are costly and lack sensitivity to detect illegitimate DNA donor integrations. In this work, we have engineered two reporter cell lines (K562_SEWAS84 and K562GWP) that efficiently quantify both the on-target and the illegitimate DNA donor integrations in a WAS-locus targeting setting. K562_SEWAS84 cells allow the detection of both HDR-and HITI-based donor integration, while K562GWP cells only report HDR-based GE. To the best of our knowledge, these are the first reporter systems that allow the use of gRNAs targeting a relevant locus to measure efficacy and specificity of DNA donor-based GE strategies. By using these models, we have found that the specificity of HDR is independent of the delivery method and that the insertion of the target sequence into the DNA donor enhances efficiency but do not affect specificity. Finally, we have also shown that the higher the number of the target sites is, the higher the specificity and efficacy of GE will be.
Keywords: CRISPR/Cas9; DNA donor; dsRED; eGFP; efficacy; homologous directed recombination (HDR); models; on-target integration; safety; specificity.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
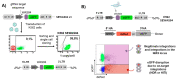





References
-
- Benabdellah K., Sanchez-Hernandez S., Aguilar-Gonzalez A., Maldonado-Perez N., Gutierrez-Guerrero A., Cortijo-Gutierrez M., Ramos-Hernandez I., Tristan-Manzano M., Galindo-Moreno P., Herrera C., et al. Genome-edited adult stem cells: Next-generation advanced therapy medicinal products (ATMPs) Stem Cells Transl. Med. 2020;9:674–685. doi: 10.1002/sctm.19-0338. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

