ASCL1 regulates neurodevelopmental transcription factors and cell cycle genes in brain tumors of glioma mouse models
- PMID: 32573857
- PMCID: PMC7587013
- DOI: 10.1002/glia.23873
ASCL1 regulates neurodevelopmental transcription factors and cell cycle genes in brain tumors of glioma mouse models
Abstract
Glioblastomas (GBMs) are incurable brain tumors with a high degree of cellular heterogeneity and genetic mutations. Transcription factors that normally regulate neural progenitors and glial development are aberrantly coexpressed in GBM, conferring cancer stem-like properties to drive tumor progression and therapeutic resistance. However, the functional role of individual transcription factors in GBMs in vivo remains elusive. Here, we demonstrate that the basic-helix-loop-helix transcription factor ASCL1 regulates transcriptional targets that are central to GBM development, including neural stem cell and glial transcription factors, oncogenic signaling molecules, chromatin modifying genes, and cell cycle and mitotic genes. We also show that the loss of ASCL1 significantly reduces the proliferation of GBMs induced in the brain of a genetically relevant glioma mouse model, resulting in extended survival times. RNA-seq analysis of mouse GBM tumors reveal that the loss of ASCL1 is associated with downregulation of cell cycle genes, illustrating an important role for ASCL1 in controlling the proliferation of GBM.
Keywords: ASCL1; brain tumor; glioma development; mouse model; transcription factor function.
© 2020 The Authors. Glia published by Wiley Periodicals, Inc.
Conflict of interest statement
The authors declare no potential conflict of interest.
Figures
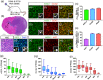


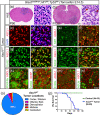
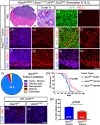

Similar articles
-
Transcription factors ASCL1 and OLIG2 drive glioblastoma initiation and co-regulate tumor cell types and migration.Nat Commun. 2024 Nov 28;15(1):10363. doi: 10.1038/s41467-024-54750-9. Nat Commun. 2024. PMID: 39609428 Free PMC article.
-
Upregulation of ASCL1 and inhibition of Notch signaling pathway characterize progressive astrocytoma.Oncogene. 2005 Oct 27;24(47):7073-83. doi: 10.1038/sj.onc.1208865. Oncogene. 2005. PMID: 16103883
-
ASCL1 Reorganizes Chromatin to Direct Neuronal Fate and Suppress Tumorigenicity of Glioblastoma Stem Cells.Cell Stem Cell. 2017 Aug 3;21(2):209-224.e7. doi: 10.1016/j.stem.2017.06.004. Epub 2017 Jul 14. Cell Stem Cell. 2017. PMID: 28712938
-
System analysis identifies distinct and common functional networks governed by transcription factor ASCL1, in glioma and small cell lung cancer.Mol Biosyst. 2017 Jul 25;13(8):1481-1494. doi: 10.1039/c6mb00851h. Mol Biosyst. 2017. PMID: 28742165
-
The Role of Neurodevelopmental Pathways in Brain Tumors.Front Cell Dev Biol. 2021 Apr 27;9:659055. doi: 10.3389/fcell.2021.659055. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34012965 Free PMC article. Review.
Cited by
-
ASCL1 Is Involved in the Pathogenesis of Schizophrenia by Regulation of Genes Related to Cell Proliferation, Neuronal Signature Formation, and Neuroplasticity.Int J Mol Sci. 2023 Oct 30;24(21):15746. doi: 10.3390/ijms242115746. Int J Mol Sci. 2023. PMID: 37958729 Free PMC article.
-
Transcription factors ASCL1 and OLIG2 drive glioblastoma initiation and co-regulate tumor cell types and migration.Nat Commun. 2024 Nov 28;15(1):10363. doi: 10.1038/s41467-024-54750-9. Nat Commun. 2024. PMID: 39609428 Free PMC article.
-
Decoding Cortical Glial Cell Development.Neurosci Bull. 2021 Apr;37(4):440-460. doi: 10.1007/s12264-021-00640-9. Epub 2021 Feb 19. Neurosci Bull. 2021. PMID: 33606177 Free PMC article.
-
GZ17-6.02 Inhibits the Growth of EGFRvIII+ Glioblastoma.Int J Mol Sci. 2022 Apr 10;23(8):4174. doi: 10.3390/ijms23084174. Int J Mol Sci. 2022. PMID: 35456993 Free PMC article.
-
Gliomas: a reflection of temporal gliogenic principles.Commun Biol. 2024 Feb 6;7(1):156. doi: 10.1038/s42003-024-05833-2. Commun Biol. 2024. PMID: 38321118 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

