Inferring generation-interval distributions from contact-tracing data
- PMID: 32574542
- PMCID: PMC7328397
- DOI: 10.1098/rsif.2019.0719
Inferring generation-interval distributions from contact-tracing data
Abstract
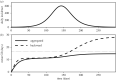
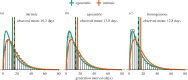
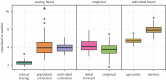
Generation intervals, defined as the time between when an individual is infected and when that individual infects another person, link two key quantities that describe an epidemic: the initial reproductive number, [Formula: see text], and the initial rate of exponential growth, r. Generation intervals can be measured through contact tracing by identifying who infected whom. We study how realized intervals differ from 'intrinsic' intervals that describe individual-level infectiousness and identify both spatial and temporal effects, including truncating (due to observation time), and the effects of susceptible depletion at various spatial scales. Early in an epidemic, we expect the variation in the realized generation intervals to be mainly driven by truncation and by the population structure near the source of disease spread; we predict that correcting realized intervals for the effect of temporal truncation but not for spatial effects will provide the initial forward generation-interval distribution, which is spatially informed and correctly links r and [Formula: see text]. We develop and test statistical methods for temporal corrections of generation intervals, and confirm our prediction using individual-based simulations on an empirical network.
Keywords: basic reproductive number; contact tracing; generation interval; infectious disease modelling; population structure.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


References
-
- Anderson RM, May RM. 1991. Infectious diseases of humans: dynamics and control. Oxford, UK: Oxford University Press.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
