Heritability and genome-wide association of swine gut microbiome features with growth and fatness parameters
- PMID: 32576852
- PMCID: PMC7311463
- DOI: 10.1038/s41598-020-66791-3
Heritability and genome-wide association of swine gut microbiome features with growth and fatness parameters
Abstract
Despite recent efforts to characterize longitudinal variation in the swine gut microbiome, the extent to which a host's genome impacts the composition of its gut microbiome is not yet well understood in pigs. The objectives of this study were: i) to identify pig gut microbiome features associated with growth and fatness, ii) to estimate the heritability of those features, and, iii) to conduct a genome-wide association study exploring the relationship between those features and single nucleotide polymorphisms (SNP) in the pig genome. A total of 1,028 pigs were characterized. Animals were genotyped with the Illumina PorcineSNP60 Beadchip. Microbiome samples from fecal swabs were obtained at weaning (Wean), at mid-test during the growth trial (MidTest), and at the end of the growth trial (OffTest). Average daily gain was calculated from birth to week 14 of the growth trial, from weaning to week 14, from week 14 to week 22, and from week 14 to harvest. Backfat and loin depth were also measured at weeks 14 and 22. Heritability estimates (±SE) of Operational Taxonomic Units ranged from 0.025 (±0.0002) to 0.139 (±0.003), from 0.029 (±0.003) to 0.289 (±0.004), and from 0.025 (±0.003) to 0.545 (±0.034) at Wean, MidTest, and OffTest, respectively. Several SNP were significantly associated with taxa at the three time points. These SNP were located in genomic regions containing a total of 68 genes. This study provides new evidence linking gut microbiome composition with growth and carcass traits in swine, while also identifying putative host genetic markers associated with significant differences in the abundance of several prevalent microbiome features.
Conflict of interest statement
The authors declare no competing interests.
Figures

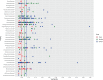
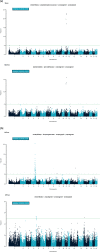

References
-
- Falony G, et al. Population-level analysis of gut microbiome variation. Science. 2016;352:560–564. - PubMed
-
- Erwin GZ, Antoon DLAK. The Host Genotype Affects the Bacterial Community in the Human Gastronintestinal Tract. Microbial Ecology in Health and Disease. 2001;13:129–134.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

