Automated CT and MRI Liver Segmentation and Biometry Using a Generalized Convolutional Neural Network
- PMID: 32582883
- PMCID: PMC7314107
- DOI: 10.1148/ryai.2019180022
Automated CT and MRI Liver Segmentation and Biometry Using a Generalized Convolutional Neural Network
Abstract
Purpose: To assess feasibility of training a convolutional neural network (CNN) to automate liver segmentation across different imaging modalities and techniques used in clinical practice and apply this to enable automation of liver biometry.
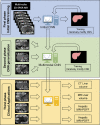
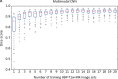
Methods: We trained a 2D U-Net CNN for liver segmentation in two stages using 330 abdominal MRI and CT exams acquired at our institution. First, we trained the neural network with non-contrast multi-echo spoiled-gradient-echo (SGPR)images with 300 MRI exams to provide multiple signal-weightings. Then, we used transfer learning to generalize the CNN with additional images from 30 contrast-enhanced MRI and CT exams.We assessed the performance of the CNN using a distinct multi-institutional data set curated from multiple sources (n = 498 subjects). Segmentation accuracy was evaluated by computing Dice scores. Utilizing these segmentations, we computed liver volume from CT and T1-weighted (T1w) MRI exams, and estimated hepatic proton- density-fat-fraction (PDFF) from multi-echo T2*w MRI exams. We compared quantitative volumetry and PDFF estimates between automated and manual segmentation using Pearson correlation and Bland-Altman statistics.
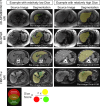
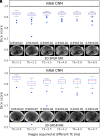
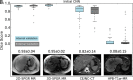
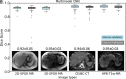
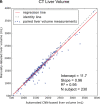
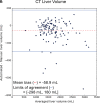
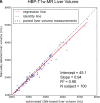
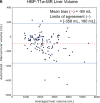
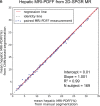
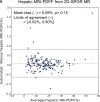
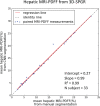
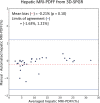
Results: Dice scores were 0.94 ± 0.06 for CT (n = 230), 0.95 ± 0.03 (n = 100) for T1w MR, and 0.92 ± 0.05 for T2*w MR (n = 169). Liver volume measured by manual and automated segmentation agreed closely for CT (95% limit-of-agreement (LoA) = [-298 mL, 180 mL]) and T1w MR (LoA = [-358 mL, 180 mL]). Hepatic PDFF measured by the two segmentations also agreed closely (LoA = [-0.62%, 0.80%]).
Conclusions: Utilizing a transfer-learning strategy, we have demonstrated the feasibility of a CNN to be generalized to perform liver segmentations across different imaging techniques and modalities. With further refinement and validation, CNNs may have broad applicability for multimodal liver volumetry and hepatic tissue characterization.
Conflict of interest statement
Figures

References
-
- Kessler LG, Barnhart HX, Buckler AJ, et al. The emerging science of quantitative imaging biomarkers terminology and definitions for scientific studies and regulatory submissions. Stat Methods Med Res 2015;24(1):9–26. - PubMed
-
- Joo S, Yang YS, Moon WK, Kim HC. Computer-aided diagnosis of solid breast nodules: use of an artificial neural network based on multiple sonographic features. IEEE Trans Med Imaging 2004;23(10):1292–1300. - PubMed
-
- Farraher SW, Jara H, Chang KJ, Hou A, Soto JA. Liver and spleen volumetry with quantitative MR imaging and dual-space clustering segmentation. Radiology 2005;237(1):322–328. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

