Programmable RNA Targeting Using CasRx in Flies
- PMID: 32584145
- PMCID: PMC7307691
- DOI: 10.1089/crispr.2020.0018
Programmable RNA Targeting Using CasRx in Flies
Erratum in
-
Correction to: Programmable RNA Targeting Using CasRx in Flies, by Buchman, et al. The CRISPR Journal 2020;3(3)164-176; doi: 10.1089/crispr.2020.0018.CRISPR J. 2024 Feb;7(1):68. doi: 10.1089/crispr.2020.0018.correx. CRISPR J. 2024. PMID: 38353625 Free PMC article. No abstract available.
Abstract
CRISPR-Cas genome editing technologies have revolutionized the fields of functional genetics and genome engineering, but with the recent discovery and optimization of RNA-targeting Cas ribonucleases, we may soon see a similar revolution in the study of RNA function and transcriptome engineering. However, to date, successful proof of principle for Cas ribonuclease RNA targeting in eukaryotic systems has been limited. Only recently has successful modification of RNA expression by a Cas ribonuclease been demonstrated in animal embryos. This previous work, however, did not evaluate endogenous expression of Cas ribonucleases and only focused on function in early developmental stages. A more comprehensive evaluation of this technology is needed to assess its potential impact. Here we report on our efforts to develop a programmable platform for RNA targeting using a Cas ribonuclease, CasRx, in the model organism Drosophila melanogaster. By genetically encoding CasRx in flies, we demonstrate moderate transcript targeting of known phenotypic genes in addition to unexpected toxicity and lethality. We also report on the off-target effects following on-target transcript cleavage by CasRx. Taken together, our results present the current state and limitations of a genetically encoded programmable RNA-targeting Cas system in Drosophila melanogaster, paving the way for future optimization of the system.
Conflict of interest statement
O.S.A and A.B. have a patent pending on this technology. All other authors declare that no competing financial interests exist.
Figures


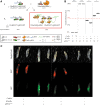
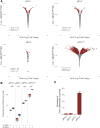
Similar articles
-
Ubiquitous and Tissue-specific RNA Targeting in Drosophila Melanogaster using CRISPR/CasRx.J Vis Exp. 2021 Feb 5;(168):10.3791/62154. doi: 10.3791/62154. J Vis Exp. 2021. PMID: 33616113 Free PMC article.
-
CRISPR-Based Technologies: Impact of RNA-Targeting Systems.Mol Cell. 2018 Nov 1;72(3):404-412. doi: 10.1016/j.molcel.2018.09.018. Mol Cell. 2018. PMID: 30388409 Free PMC article. Review.
-
Molecular Mechanisms of RNA Targeting by Cas13-containing Type VI CRISPR-Cas Systems.J Mol Biol. 2019 Jan 4;431(1):66-87. doi: 10.1016/j.jmb.2018.06.029. Epub 2018 Jun 22. J Mol Biol. 2019. PMID: 29940185 Review.
-
Efficient multiplexed genome engineering with a polycistronic tRNA and CRISPR guide-RNA reveals an important role of detonator in reproduction of Drosophila melanogaster.PLoS One. 2021 Jan 14;16(1):e0245454. doi: 10.1371/journal.pone.0245454. eCollection 2021. PLoS One. 2021. PMID: 33444382 Free PMC article.
-
Efficient Screening of CRISPR/Cas9-Induced Events in Drosophila Using a Co-CRISPR Strategy.G3 (Bethesda). 2017 Jan 5;7(1):87-93. doi: 10.1534/g3.116.036723. G3 (Bethesda). 2017. PMID: 27793971 Free PMC article.
Cited by
-
A target expression threshold dictates invader defense and prevents autoimmunity by CRISPR-Cas13.Cell Host Microbe. 2022 Aug 10;30(8):1151-1162.e6. doi: 10.1016/j.chom.2022.05.013. Epub 2022 Jun 10. Cell Host Microbe. 2022. PMID: 35690065 Free PMC article.
-
Enhanced RNA-targeting CRISPR-Cas technology in zebrafish.Nat Commun. 2025 Mar 16;16(1):2591. doi: 10.1038/s41467-025-57792-9. Nat Commun. 2025. PMID: 40091120 Free PMC article.
-
Massively Parallel Profiling of RNA-targeting CRISPR-Cas13d.bioRxiv [Preprint]. 2023 Mar 28:2023.03.27.534188. doi: 10.1101/2023.03.27.534188. bioRxiv. 2023. Update in: Nat Commun. 2024 Jan 12;15(1):498. doi: 10.1038/s41467-024-44738-w. PMID: 37034598 Free PMC article. Updated. Preprint.
-
dCas13-mediated translational repression for accurate gene silencing in mammalian cells.Nat Commun. 2024 Mar 11;15(1):2205. doi: 10.1038/s41467-024-46412-7. Nat Commun. 2024. PMID: 38467613 Free PMC article.
-
Precise genome engineering in Drosophila using prime editing.Proc Natl Acad Sci U S A. 2021 Jan 5;118(1):e2021996118. doi: 10.1073/pnas.2021996118. Proc Natl Acad Sci U S A. 2021. PMID: 33443210 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
