A Single-Neuron: Current Trends and Future Prospects
- PMID: 32585883
- PMCID: PMC7349798
- DOI: 10.3390/cells9061528
A Single-Neuron: Current Trends and Future Prospects
Abstract
The brain is an intricate network with complex organizational principles facilitating a concerted communication between single-neurons, distinct neuron populations, and remote brain areas. The communication, technically referred to as connectivity, between single-neurons, is the center of many investigations aimed at elucidating pathophysiology, anatomical differences, and structural and functional features. In comparison with bulk analysis, single-neuron analysis can provide precise information about neurons or even sub-neuron level electrophysiology, anatomical differences, pathophysiology, structural and functional features, in addition to their communications with other neurons, and can promote essential information to understand the brain and its activity. This review highlights various single-neuron models and their behaviors, followed by different analysis methods. Again, to elucidate cellular dynamics in terms of electrophysiology at the single-neuron level, we emphasize in detail the role of single-neuron mapping and electrophysiological recording. We also elaborate on the recent development of single-neuron isolation, manipulation, and therapeutic progress using advanced micro/nanofluidic devices, as well as microinjection, electroporation, microelectrode array, optical transfection, optogenetic techniques. Further, the development in the field of artificial intelligence in relation to single-neurons is highlighted. The review concludes with between limitations and future prospects of single-neuron analyses.
Keywords: artificial intelligence; electrophysiological recording; isolation; mapping; micro/nanofluidic devices; microelectrode array; single-neuron models; therapy; transfection.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

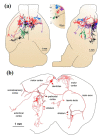
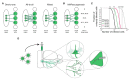
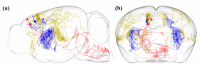
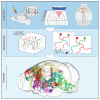




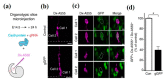

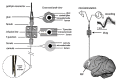
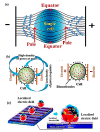
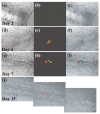
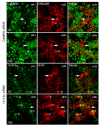
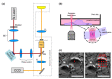
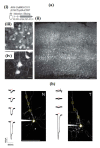
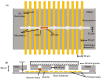

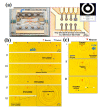
References
-
- Santra T.S., Tseng F.-G. Handbook of Single Cell Technologies. Springer Nature: Singapore Pte. Ltd; Singapore: 2018.
-
- Santra T.S., Tseng F.-G. Micro/Nanofluidic Devices for Single Cell Analysis. J. Micromach. 2014;5:154–157. doi: 10.3390/mi5020154. - DOI
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

