Mining ancient microbiomes using selective enrichment of damaged DNA molecules
- PMID: 32586278
- PMCID: PMC7318760
- DOI: 10.1186/s12864-020-06820-7
Mining ancient microbiomes using selective enrichment of damaged DNA molecules
Abstract
Background: The identification of bona fide microbial taxa in microbiomes derived from ancient and historical samples is complicated by the unavoidable mixture between DNA from ante- and post-mortem microbial colonizers. One possibility to distinguish between these sources of microbial DNA is querying for the presence of age-associated degradation patterns typical of ancient DNA (aDNA). The presence of uracils, resulting from cytosine deamination, has been detected ubiquitously in aDNA retrieved from diverse sources, and used as an authentication criterion. Here, we employ a library preparation method that separates molecules that carry uracils from those that do not for a set of samples that includes Neandertal remains, herbarium specimens and archaeological plant remains.
Results: We show that sequencing DNA libraries enriched in molecules carrying uracils effectively amplifies age associated degradation patterns in microbial mixtures of ancient and historical origin. This facilitates the discovery of authentic ancient microbial taxa in cases where degradation patterns are difficult to detect due to large sequence divergence in microbial mixtures. Additionally, the relative enrichment of taxa in the uracil enriched fraction can help to identify bona fide ancient microbial taxa that could be missed using a more targeted approach.
Conclusions: Our experiments show, that in addition to its use in enriching authentic endogenous DNA of organisms of interest, the selective enrichment of damaged DNA molecules can be a valuable tool in the discovery of ancient microbial taxa.
Keywords: Ancient DNA; Authentication; Metagenomics.
Conflict of interest statement
The authors declare no competing interests.
Figures
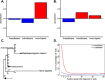


References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

