This is a preprint.
The D614G mutation in the SARS-CoV-2 spike protein reduces S1 shedding and increases infectivity
- PMID: 32587973
- PMCID: PMC7310631
- DOI: 10.1101/2020.06.12.148726
The D614G mutation in the SARS-CoV-2 spike protein reduces S1 shedding and increases infectivity
Update in
-
SARS-CoV-2 spike-protein D614G mutation increases virion spike density and infectivity.Nat Commun. 2020 Nov 26;11(1):6013. doi: 10.1038/s41467-020-19808-4. Nat Commun. 2020. PMID: 33243994 Free PMC article.
Abstract
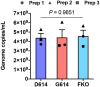
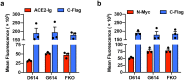
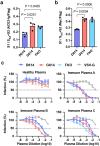
SARS coronavirus 2 (SARS-CoV-2) isolates encoding a D614G mutation in the viral spike (S) protein predominate over time in locales where it is found, implying that this change enhances viral transmission. We therefore compared the functional properties of the S proteins with aspartic acid (S D614 ) and glycine (S G614 ) at residue 614. We observed that retroviruses pseudotyped with S G614 infected ACE2-expressing cells markedly more efficiently than those with S D614 . This greater infectivity was correlated with less S1 shedding and greater incorporation of the S protein into the pseudovirion. Similar results were obtained using the virus-like particles produced with SARS-CoV-2 M, N, E, and S proteins. However, S G614 did not bind ACE2 more efficiently than S D614 , and the pseudoviruses containing these S proteins were neutralized with comparable efficiencies by convalescent plasma. These results show S G614 is more stable than S D614 , consistent with epidemiological data suggesting that viruses with S G614 transmit more efficiently.
Conflict of interest statement
Competing interests
The authors claim no competing interest.
Figures




References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
