Structural and Chemical Biology of the Interaction of Cyclooxygenase with Substrates and Non-Steroidal Anti-Inflammatory Drugs
- PMID: 32609495
- PMCID: PMC8253488
- DOI: 10.1021/acs.chemrev.0c00215
Structural and Chemical Biology of the Interaction of Cyclooxygenase with Substrates and Non-Steroidal Anti-Inflammatory Drugs
Abstract
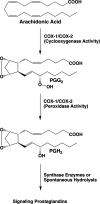
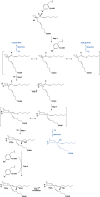
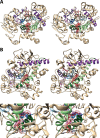
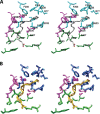
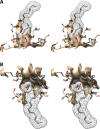
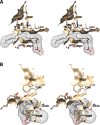
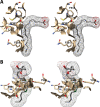
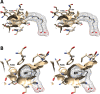
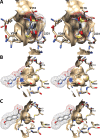
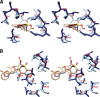
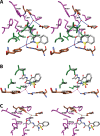
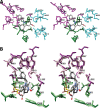
Cyclooxgenases are key enzymes of lipid signaling. They carry out the first step in the production of prostaglandins, important mediators of inflammation, pain, cardiovascular disease, and cancer, and they are the molecular targets for nonsteroidal anti-inflammatory drugs, which are among the oldest and most chemically diverse set of drugs known. Homodimeric proteins that behave as allosterically modulated, functional heterodimers, the cyclooxygenases exhibit complex kinetic behavior, requiring peroxide-dependent activation and undergoing suicide inactivation. Due to their important physiological and pathophysiological roles and keen interest on the part of the pharmaceutical industry, the cyclooxygenases have been the focus of a vast array of structural studies, leading to the publication of over 80 crystal structures of the enzymes in complex with substrates or inhibitors supported by a wealth of functional data generated by site-directed mutation experiments. In this review, we explore the chemical biology of the cyclooxygenases through the lens of this wealth of structural and functional information. We identify key structural features of the cyclooxygenases, break down their active site into regional binding pockets to facilitate comparisons between structures, and explore similarities and differences in the binding modes of the wide variety of ligands (both substrates and inhibitors) that have been characterized in complex with the enzymes. Throughout, we correlate structure with function whenever possible. Finally, we summarize what can and cannot be learned from the currently available structural data and discuss the critical intriguing questions that remain despite the wealth of information that has been amassed in this field.
Conflict of interest statement
The authors declare no competing financial interest.
Figures






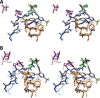
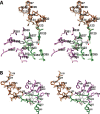
References
-
- Smith W. Molecular Biology of Prostanoid Biosynthetic Enzymes and Receptors. Adv. Exp. Med. Biol. 1997, 400B, 989–1011. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

