Implementing cell-free DNA of pancreatic cancer patient-derived organoids for personalized oncology
- PMID: 32614802
- PMCID: PMC7455062
- DOI: 10.1172/jci.insight.137809
Implementing cell-free DNA of pancreatic cancer patient-derived organoids for personalized oncology
Abstract
One of the major challenges in using pancreatic cancer patient-derived organoids (PDOs) in precision oncology is the time from biopsy to functional characterization. This is particularly true for endoscopic ultrasound-guided fine-needle aspiration biopsies, typically resulting in specimens with limited tumor cell yield. Here, we tested conditioned media of individual PDOs for cell-free DNA to detect driver mutations already early on during the expansion process to accelerate the genetic characterization of PDOs as well as subsequent functional testing. Importantly, genetic alterations detected in the PDO supernatant, collected as early as 72 hours after biopsy, recapitulate the mutational profile of the primary tumor, indicating suitability of this approach to subject PDOs to drug testing in a reduced time frame. In addition, we demonstrated that this workflow was practicable, even in patients for whom the amount of tumor material was not sufficient for molecular characterization by established means. Together, our findings demonstrate that generating PDOs from very limited biopsy material permits molecular profiling and drug testing. With our approach, this can be achieved in a rapid and feasible fashion with broad implications in clinical practice.
Keywords: Gastroenterology; Oncology; Translation.
Conflict of interest statement
Figures
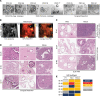
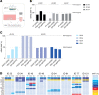
Similar articles
-
Development of Patient-Derived Gastric Cancer Organoids from Endoscopic Biopsies and Surgical Tissues.Ann Surg Oncol. 2018 Sep;25(9):2767-2775. doi: 10.1245/s10434-018-6662-8. Epub 2018 Jul 12. Ann Surg Oncol. 2018. PMID: 30003451
-
Organoids for Functional Precision Medicine in Advanced Pancreatic Cancer.Gastroenterology. 2024 Oct;167(5):961-976.e13. doi: 10.1053/j.gastro.2024.05.032. Epub 2024 Jun 10. Gastroenterology. 2024. PMID: 38866343
-
Implementing patient derived organoids in functional precision medicine for patients with advanced colorectal cancer.J Exp Clin Cancer Res. 2023 Oct 25;42(1):281. doi: 10.1186/s13046-023-02853-4. J Exp Clin Cancer Res. 2023. PMID: 37880806 Free PMC article.
-
Patient derived organoids in prostate cancer: improving therapeutic efficacy in precision medicine.Mol Cancer. 2021 Sep 29;20(1):125. doi: 10.1186/s12943-021-01426-3. Mol Cancer. 2021. PMID: 34587953 Free PMC article. Review.
-
The use of patient-derived xenografts and patient-derived organoids in the search for new therapeutic regimens for pancreatic carcinoma. A review.Biomed Pharmacother. 2025 Jan;182:117750. doi: 10.1016/j.biopha.2024.117750. Epub 2024 Dec 16. Biomed Pharmacother. 2025. PMID: 39689516 Review.
Cited by
-
The functional and clinical roles of liquid biopsy in patient-derived models.J Hematol Oncol. 2023 Apr 8;16(1):36. doi: 10.1186/s13045-023-01433-5. J Hematol Oncol. 2023. PMID: 37031172 Free PMC article. Review.
-
Musculoskeletal biomarkers in health and disease: implications for the aging spine-a review for spinal surgeons by the SRS adult spinal deformity task force on senescence.Spine Deform. 2025 Jun 4. doi: 10.1007/s43390-025-01124-w. Online ahead of print. Spine Deform. 2025. PMID: 40465097 Review.
-
The pivotal application of patient-derived organoid biobanks for personalized treatment of gastrointestinal cancers.Biomark Res. 2022 Oct 8;10(1):73. doi: 10.1186/s40364-022-00421-0. Biomark Res. 2022. PMID: 36207749 Free PMC article. Review.
-
Spatiotemporal dynamics of self-organized branching in pancreas-derived organoids.Nat Commun. 2022 Sep 5;13(1):5219. doi: 10.1038/s41467-022-32806-y. Nat Commun. 2022. PMID: 36064947 Free PMC article.
-
Inspiring a convergent engineering approach to measure and model the tissue microenvironment.Heliyon. 2024 Jun 8;10(12):e32546. doi: 10.1016/j.heliyon.2024.e32546. eCollection 2024 Jun 30. Heliyon. 2024. PMID: 38975228 Free PMC article. Review.
References
-
- Aguirre AJ, et al. Real-time genomic characterization of advanced pancreatic cancer to enable precision medicine. Cancer Discov. 2018;8(9):1096–1111. doi: 10.1158/2159-8290.CD-18-0275. - DOI - PMC - PubMed
-
- Tiriac H, et al. Organoid profiling identifies common responders to chemotherapy in pancreatic cancer. Cancer Discov. 2018;8(9):1112–1129. doi: 10.1158/2159-8290.CD-18-0349. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

