Fine-scale succession patterns and assembly mechanisms of bacterial community of Litopenaeus vannamei larvae across the developmental cycle
- PMID: 32620132
- PMCID: PMC7334860
- DOI: 10.1186/s40168-020-00879-w
Fine-scale succession patterns and assembly mechanisms of bacterial community of Litopenaeus vannamei larvae across the developmental cycle
Abstract
Background: Microbiome assembly in early life may have a long-term impact on host health. Larval nursery is a crucial period that determines the success in culture of Litopenaeus vannamei, the most productive shrimp species in world aquaculture industry. However, the succession patterns and assembly mechanisms of larval shrimp bacterial community still lack characterization at a fine temporal scale. Here, using a high-frequency sampling strategy and 16S rRNA gene amplicon sequencing, we investigated dynamics of larval shrimp bacterial community and its relationship with bacterioplankton in the rearing water across the whole developmental cycle in a realistic aquaculture practice.
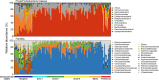
Results: Alpha-diversity of larval shrimp bacteria showed a U-shaped pattern across the developmental cycle with the stages zoea and mysis as the valley. Correspondingly, the compositions of dominant bacterial taxa at the stages nauplius and early postlarvae were more complex than other stages. Remarkably, Rhodobacteraceae maintained the overwhelming dominance after the mouth opening of larvae (zoea I~early postlarvae). The taxonomic and phylogenetic compositions of larval bacterial community both showed stage-dependent patterns with higher rate of taxonomic turnover, suggesting that taxonomic turnover was mainly driven by temporal switching among closely related taxa (such as Rhodobacteraceae taxa). The assembly of larval bacteria was overall governed by neutral processes (dispersal among individuals and ecological drift) at all the stages, but bacterioplankton also had certain contribution during three sub-stages of zoea, when larval and water bacterial communities were most associated. Furthermore, the positive host selection for Rhodobacteraceae taxa from the rearing water during the zoea stage and its persistent dominance and large predicted contribution to metabolic potentials of organic matters at post-mouth opening stages suggest a crucial role of this family in larval microbiome and thus a potential source of probiotic candidates for shrimp larval nursery.
Conclusions: Our results reveal pronounced succession patterns and dynamic assembly processes of larval shrimp bacterial communities during the developmental cycle, highlighting the importance of the mouth opening stage from the perspective of microbial ecology. We also suggest the possibility and potential timing in microbial management of the rearing water for achieving the beneficial larval microbiota in the nursery practice. Video Abstract.
Keywords: Community assembly; Early life microbiome; Host development; Shrimp larvae; Succession pattern.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





References
-
- Matamoros S, Gras-Leguen C, Le Vacon F, Potel G, de La Cochetiere MF. Development of intestinal microbiota in infants and its impact on health. Trends Microbiol. 2013;21:167–173. - PubMed
-
- Bárcena C, Valdés-Mas R, Mayoral P, Garabaya C, Durand S, Rodríguez F, et al. Healthspan and lifespan extension by fecal microbiota transplantation into progeroid mice. Nat Med. 2019;25:1234–1242. - PubMed
-
- Zokaeifar H, Luis Balcazar J, Saad CR, Kamarudin MS, Sijam K, Arshad A, et al. Effects of Bacillus subtilis on the growth performance, digestive enzymes, immune gene expression and disease resistance of white shrimp, Litopenaeus vannamei. Fish Shellfish Immunol. 2012;33:683–689. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- LY18C030002/Zhejiang Provincial Natural Science Foundation of China/International
- 31672658/National Natural Science Foundation of China/International
- 2017C110001/Agricultural Major Project of Ningbo, China/International
- LGN20C190008/Basic Public Welfare Research Project of Zhejiang Province/International
LinkOut - more resources
Full Text Sources

