Multimodal single-cell analysis reveals distinct radioresistant stem-like and progenitor cell populations in murine glioma
- PMID: 32621641
- PMCID: PMC7586969
- DOI: 10.1002/glia.23866
Multimodal single-cell analysis reveals distinct radioresistant stem-like and progenitor cell populations in murine glioma
Abstract
Radiation therapy is part of the standard of care for gliomas and kills a subset of tumor cells, while also altering the tumor microenvironment. Tumor cells with stem-like properties preferentially survive radiation and give rise to glioma recurrence. Various techniques for enriching and quantifying cells with stem-like properties have been used, including the fluorescence activated cell sorting (FACS)-based side population (SP) assay, which is a functional assay that enriches for stem-like tumor cells. In these analyses, mouse models of glioma have been used to understand the biology of this disease and therapeutic responses, including the radiation response. We present combined SP analysis and single-cell RNA sequencing of genetically-engineered mouse models of glioma to show a time course of cellular response to radiation. We identify and characterize two distinct tumor cell populations that are inherently radioresistant and also distinct effects of radiation on immune cell populations within the tumor microenvironment.
Keywords: SP analysis; glioma; glioma stem cells; myeloid cells; radiation response; radioresistance; single-cell RNA sequencing.
© 2020 The Authors. Glia published by Wiley Periodicals LLC.
Figures
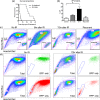





References
-
- Bao, S. , Wu, Q. , McLendon, R. E. , Hao, Y. , Shi, Q. , Hjelmeland, A. B. , … Rich, J. N. (2006). Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature, 444(7120), 756–760. - PubMed
-
- Bennett, M. L. , Chris Bennett, F. , Liddelow, S. A. , Ajami, B. , Zamanian, J. L. , Fernhoff, N. B. , … Barres, B. A. (2016). New tools for studying microglia in the mouse and human CNS. Proceedings of the National Academy of Sciences, 113(12), E1738–E1746. 10.1073/pnas.1525528113 - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

