Genome wide distribution of G-quadruplexes and their impact on gene expression in malaria parasites
- PMID: 32628663
- PMCID: PMC7365481
- DOI: 10.1371/journal.pgen.1008917
Genome wide distribution of G-quadruplexes and their impact on gene expression in malaria parasites
Abstract
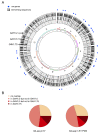
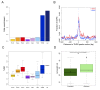
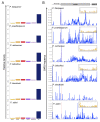
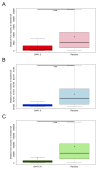
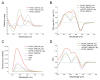
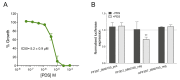
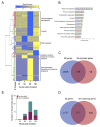
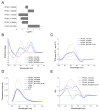
Mechanisms of transcriptional control in malaria parasites are still not fully understood. The positioning patterns of G-quadruplex (G4) DNA motifs in the parasite's AT-rich genome, especially within the var gene family which encodes virulence factors, and in the vicinity of recombination hotspots, points towards a possible regulatory role of G4 in gene expression and genome stability. Here, we carried out the most comprehensive genome-wide survey, to date, of G4s in the Plasmodium falciparum genome using G4Hunter, which identifies G4 forming sequences (G4FS) considering their G-richness and G-skewness. We show an enrichment of G4FS in nucleosome-depleted regions and in the first exon of var genes, a pattern that is conserved within the closely related Laverania Plasmodium parasites. Under G4-stabilizing conditions, i.e., following treatment with pyridostatin (a high affinity G4 ligand), we show that a bona fide G4 found in the non-coding strand of var promoters modulates reporter gene expression. Furthermore, transcriptional profiling of pyridostatin-treated parasites, shows large scale perturbations, with deregulation affecting for instance the ApiAP2 family of transcription factors and genes involved in ribosome biogenesis. Overall, our study highlights G4s as important DNA secondary structures with a role in Plasmodium gene expression regulation, sub-telomeric recombination and var gene biology.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

References
-
- WHO. World malaria report 2018. WHO website. 2018. http://www.who.int/malaria/publications/world-malaria-report-2018/report...
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

