Using unnatural amino acids to selectively label proteins for cellular imaging: a cell biologist viewpoint
- PMID: 32640070
- PMCID: PMC7983921
- DOI: 10.1111/febs.15477
Using unnatural amino acids to selectively label proteins for cellular imaging: a cell biologist viewpoint
Abstract
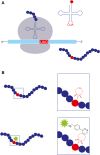
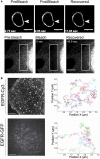
Twenty-five years ago, GFP revolutionized the field of cell biology by enabling scientists to visualize, for the first time, proteins in living cells. However, when it comes to current, state-of-the-art imaging technologies, fluorescent proteins (such as GFP) have several limitations that result from their size and photophysics. Over the past decade, an elegant, alternative approach, which is based on the direct labeling of proteins with fluorescent dyes and is compatible with live-cell and super-resolution imaging applications, has been introduced. In this approach, an unnatural amino acid that can covalently bind a fluorescent dye is incorporated into the coding sequence of a protein. The protein of interest is thereby site-specifically fluorescently labeled inside the cell, eliminating the need for protein- or peptide-labeling tags. Whether this labeling approach will change cell biology research is currently unclear, but it clearly has the potential to do so. In this short review, a general overview of this approach is provided, focusing on the imaging of site-specifically labeled proteins in mammalian tissue culture cells, and highlighting its advantages and limitations for cellular imaging.
Keywords: bioorthogonal reactions; click chemistry; fluorescent dyes; genetic code expansion; light microscopy; noncanonical amino acids; protein labeling.
© 2020 The Authors. The FEBS Journal published by John Wiley & Sons Ltd on behalf of Federation of European Biochemical Societies.
Conflict of interest statement
The author declare no conflict of interest.
Figures

Similar articles
-
Genetic Code Expansion- and Click Chemistry-Based Site-Specific Protein Labeling for Intracellular DNA-PAINT Imaging.Methods Mol Biol. 2018;1728:279-295. doi: 10.1007/978-1-4939-7574-7_18. Methods Mol Biol. 2018. PMID: 29405005
-
Bioorthogonal Click Chemistry Enables Site-specific Fluorescence Labeling of Functional NMDA Receptors for Super-Resolution Imaging.Angew Chem Int Ed Engl. 2018 Dec 10;57(50):16364-16369. doi: 10.1002/anie.201808951. Epub 2018 Nov 15. Angew Chem Int Ed Engl. 2018. PMID: 30347512
-
Site-Specific Bioorthogonal Labeling for Fluorescence Imaging of Intracellular Proteins in Living Cells.J Am Chem Soc. 2016 Nov 2;138(43):14423-14433. doi: 10.1021/jacs.6b08733. Epub 2016 Oct 21. J Am Chem Soc. 2016. PMID: 27768298 Free PMC article.
-
Chemical biology-based approaches on fluorescent labeling of proteins in live cells.Mol Biosyst. 2013 May;9(5):862-72. doi: 10.1039/c2mb25422k. Mol Biosyst. 2013. PMID: 23318293 Review.
-
Chemical tags for site-specific fluorescent labeling of biomolecules.Amino Acids. 2016 Jun;48(6):1357-72. doi: 10.1007/s00726-016-2204-5. Epub 2016 Mar 11. Amino Acids. 2016. PMID: 26969255 Review.
Cited by
-
Insights into receptor structure and dynamics at the surface of living cells.Nat Commun. 2023 Mar 22;14(1):1596. doi: 10.1038/s41467-023-37284-4. Nat Commun. 2023. PMID: 36949079 Free PMC article.
-
New Amino Acid-Based Thiosemicarbazones and Hydrazones: Synthesis and Evaluation as Fluorimetric Chemosensors in Aqueous Mixtures.Molecules. 2023 Oct 25;28(21):7256. doi: 10.3390/molecules28217256. Molecules. 2023. PMID: 37959675 Free PMC article.
-
Optimized genetic code expansion technology for time-dependent induction of adhesion GPCR-ligand engagement.Protein Sci. 2023 Apr;32(4):e4614. doi: 10.1002/pro.4614. Protein Sci. 2023. PMID: 36870000 Free PMC article.
-
Applications of genetic code expansion technology in eukaryotes.Protein Cell. 2024 May 7;15(5):331-363. doi: 10.1093/procel/pwad051. Protein Cell. 2024. PMID: 37847216 Free PMC article. Review.
-
Bioorthogonal Reactions in Bioimaging.Top Curr Chem (Cham). 2024 Feb 24;382(1):7. doi: 10.1007/s41061-024-00452-1. Top Curr Chem (Cham). 2024. PMID: 38400853 Free PMC article. Review.
References
-
- Lang K & Chin JW (2014) Cellular incorporation of unnatural amino acids and bioorthogonal labeling of proteins. Chem Rev 114, 4764–4806. - PubMed
-
- Nikic I & Lemke EA (2015) Genetic code expansion enabled site‐specific dual‐color protein labeling: superresolution microscopy and beyond. Curr Opin Chem Biol 28, 164–173. - PubMed
-
- Beatty KE, Liu JC, Xie F, Dieterich DC, Schuman EM, Wang Q & Tirrell DA (2006) Fluorescence visualization of newly synthesized proteins in mammalian cells. Angew Chem Int Ed Engl 45, 7364–7367. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

