Observing Protein Degradation by the PAN-20S Proteasome by Time-Resolved Neutron Scattering
- PMID: 32640186
- PMCID: PMC7376118
- DOI: 10.1016/j.bpj.2020.06.015
Observing Protein Degradation by the PAN-20S Proteasome by Time-Resolved Neutron Scattering
Abstract
The proteasome is a key player of regulated protein degradation in all kingdoms of life. Although recent atomic structures have provided snapshots on a number of conformations, data on substrate states and populations during the active degradation process in solution remain scarce. Here, we use time-resolved small-angle neutron scattering of a deuterium-labeled GFPssrA substrate and an unlabeled archaeal PAN-20S system to obtain direct structural information on substrate states during ATP-driven unfolding and subsequent proteolysis in solution. We find that native GFPssrA structures are degraded in a biexponential process, which correlates strongly with ATP hydrolysis, the loss of fluorescence, and the buildup of small oligopeptide products. Our solution structural data support a model in which the substrate is directly translocated from PAN into the 20S proteolytic chamber, after a first, to our knowledge, successful unfolding process that represents a point of no return and thus prevents dissociation of the complex and the release of harmful, aggregation-prone products.
Copyright © 2020 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures
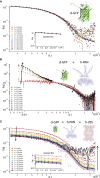




Comment in
-
Hiding the Elephant in the Room with Experimental Neutrons.Biophys J. 2020 Jul 21;119(2):234-235. doi: 10.1016/j.bpj.2020.05.038. Epub 2020 Jun 24. Biophys J. 2020. PMID: 32640187 Free PMC article. No abstract available.
References
-
- Wolf D.H., Menssen R. Mechanisms of cell regulation - proteolysis, the big surprise. FEBS Lett. 2018;592:2515–2524. - PubMed
-
- Alberts B., Johnson A., Walter P. Garland Science; Adingdon, NY: 2008. Molecular Biology of the Cell.
-
- Goldberg A.L. Protein degradation and protection against misfolded or damaged proteins. Nature. 2003;426:895–899. - PubMed
-
- Schoenheimer R. Harvard University Press; Cambridge, MA: 1946. The Dynamic State of Body Constituents.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

