Rhizobium leguminosarum bv. trifolii NodD2 Enhances Competitive Nodule Colonization in the Clover-Rhizobium Symbiosis
- PMID: 32651206
- PMCID: PMC7480365
- DOI: 10.1128/AEM.01268-20
Rhizobium leguminosarum bv. trifolii NodD2 Enhances Competitive Nodule Colonization in the Clover-Rhizobium Symbiosis
Abstract
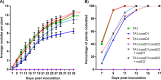
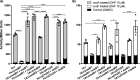
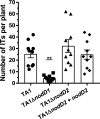
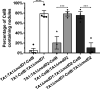
Establishment of the symbiotic relationship that develops between rhizobia and their legume hosts is contingent upon an interkingdom signal exchange. In response to host legume flavonoids, NodD proteins from compatible rhizobia activate expression of nodulation genes that produce lipochitin oligosaccharide signaling molecules known as Nod factors. Root nodule formation commences upon legume recognition of compatible Nod factor. Rhizobium leguminosarum was previously considered to contain one copy of nodD; here, we show that some strains of the Trifolium (clover) microsymbiont R. leguminosarum bv. trifolii contain a second copy designated nodD2. nodD2 genes were present in 8 out of 13 strains of R. leguminosarum bv. trifolii, but were absent from the genomes of 16 R. leguminosarum bv. viciae strains. Analysis of single and double nodD1 and nodD2 mutants in R. leguminosarum bv. trifolii strain TA1 revealed that NodD2 was functional and enhanced nodule colonization competitiveness. However, NodD1 showed significantly greater capacity to induce nod gene expression and infection thread formation. Clover species are either annual or perennial and this phenological distinction is rarely crossed by individual R. leguminosarum bv. trifolii microsymbionts for effective symbiosis. Of 13 strains with genome sequences available, 7 of the 8 effective microsymbionts of perennial hosts contained nodD2, whereas the 3 microsymbionts of annual hosts did not. We hypothesize that NodD2 inducer recognition differs from NodD1, and NodD2 functions to enhance competition and effective symbiosis, which may discriminate in favor of perennial hosts.IMPORTANCE Establishment of the rhizobium-legume symbiosis requires a highly specific and complex signal exchange between both participants. Rhizobia perceive legume flavonoid compounds through LysR-type NodD regulators. Often, rhizobia encode multiple copies of nodD, which is one determinant of host specificity. In some species of rhizobia, the presence of multiple copies of NodD extends their symbiotic host-range. Here, we identified and characterized a second copy of nodD present in some strains of the clover microsymbiont Rhizobium leguminosarum bv. trifolii. The second nodD gene contributed to the competitive ability of the strain on white clover, an important forage legume. A screen for strains containing nodD2 could be utilized as one criterion to select strains with enhanced competitive ability for use as inoculants for pasture production.
Keywords: Nod factor; NodD; Rhizobium leguminosarum; clover; competition; symbiosis.
Copyright © 2020 American Society for Microbiology.
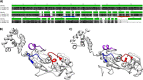
Figures

Similar articles
-
Response to flavonoids as a factor influencing competitiveness and symbiotic activity of Rhizobium leguminosarum.Microbiol Res. 2010;165(1):50-60. doi: 10.1016/j.micres.2008.06.002. Epub 2008 Aug 3. Microbiol Res. 2010. PMID: 18678476
-
Rhizobium leguminosarum bv. trifolii rosR is required for interaction with clover, biofilm formation and adaptation to the environment.BMC Microbiol. 2010 Nov 11;10:284. doi: 10.1186/1471-2180-10-284. BMC Microbiol. 2010. PMID: 21070666 Free PMC article.
-
Rhizobium meliloti has three functional copies of the nodD symbiotic regulatory gene.Proc Natl Acad Sci U S A. 1987 Dec;84(23):8558-62. doi: 10.1073/pnas.84.23.8558. Proc Natl Acad Sci U S A. 1987. PMID: 3479806 Free PMC article.
-
Specificity in Legume-Rhizobia Symbioses.Int J Mol Sci. 2017 Mar 26;18(4):705. doi: 10.3390/ijms18040705. Int J Mol Sci. 2017. PMID: 28346361 Free PMC article. Review.
-
Quorum sensing in nitrogen-fixing rhizobia.Microbiol Mol Biol Rev. 2003 Dec;67(4):574-92. doi: 10.1128/MMBR.67.4.574-592.2003. Microbiol Mol Biol Rev. 2003. PMID: 14665677 Free PMC article. Review.
Cited by
-
Quantitative genetic analysis reveals potential to breed for improved white clover growth in symbiosis with nitrogen-fixing Rhizobium bacteria.Front Plant Sci. 2022 Sep 20;13:953400. doi: 10.3389/fpls.2022.953400. eCollection 2022. Front Plant Sci. 2022. PMID: 36212301 Free PMC article.
-
Legume-rhizobium dance: an agricultural tool that could be improved?Microb Biotechnol. 2021 Sep;14(5):1897-1917. doi: 10.1111/1751-7915.13906. Epub 2021 Jul 28. Microb Biotechnol. 2021. PMID: 34318611 Free PMC article. Review.
-
Flavonoids affect the endophytic bacterial community in Ginkgo biloba leaves with increasing altitude.Front Plant Sci. 2022 Aug 11;13:982771. doi: 10.3389/fpls.2022.982771. eCollection 2022. Front Plant Sci. 2022. PMID: 36035669 Free PMC article.
-
What determines symbiotic nitrogen fixation efficiency in rhizobium: recent insights into Rhizobium leguminosarum.Arch Microbiol. 2023 Aug 5;205(9):300. doi: 10.1007/s00203-023-03640-7. Arch Microbiol. 2023. PMID: 37542687 Review.
-
Adaptive Evolution of Rhizobial Symbiosis beyond Horizontal Gene Transfer: From Genome Innovation to Regulation Reconstruction.Genes (Basel). 2023 Jan 20;14(2):274. doi: 10.3390/genes14020274. Genes (Basel). 2023. PMID: 36833201 Free PMC article. Review.
References
-
- Hynes MF, Finan TM. 1998. General genetic knowledge, p 25–43, The Rhizobiaceae. Springer.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

