Epidemics as an adaptive driving force determining lifespan setpoints
- PMID: 32651271
- PMCID: PMC7395509
- DOI: 10.1073/pnas.1920988117
Epidemics as an adaptive driving force determining lifespan setpoints
Abstract
Species-specific limits to lifespan (lifespan setpoint) determine the life expectancy of any given organism. Whether limiting lifespan provides an evolutionary benefit or is the result of an inevitable decline in fitness remains controversial. The identification of mutations extending lifespan suggests that aging is under genetic control, but the evolutionary driving forces limiting lifespan have not been defined. By examining the impact of lifespan on pathogen spread in a population, we propose that epidemics drive lifespan setpoints' evolution. Shorter lifespan limits infection spread and accelerates pathogen clearance when compared to populations with longer-lived individuals. Limiting longevity is particularly beneficial in the context of zoonotic transmissions, where pathogens must undergo adaptation to a new host. Strikingly, in populations exposed to pathogens, shorter-living variants outcompete individuals with longer lifespans. We submit that infection outbreaks can contribute to control the evolution of species' lifespan setpoints.
Keywords: aging; epidemics; evolution; lifespan.
Copyright © 2020 the Author(s). Published by PNAS.
Conflict of interest statement
The authors declare no competing interest.
Figures



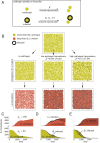
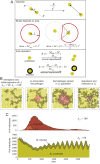


References
-
- Medawar P. B., An Unsolved Problem of Biology (H. K. Lewis, London, 1952).
-
- Kirkwood T. B., Holliday R., The evolution of ageing and longevity. Proc. R. Soc. Lond. B Biol. Sci. 205, 531–546 (1979). - PubMed
-
- Williams G. C., Pleiotropy, natural selection, and the evolution of senescence. Evolution 11, 398–411 (1957).

