KCNQs: Ligand- and Voltage-Gated Potassium Channels
- PMID: 32655402
- PMCID: PMC7324551
- DOI: 10.3389/fphys.2020.00583
KCNQs: Ligand- and Voltage-Gated Potassium Channels
Abstract
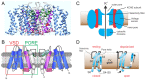
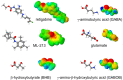
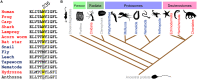
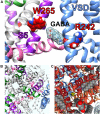
Voltage-gated potassium (Kv) channels in the KCNQ (Kv7) family are essential features of a broad range of excitable and non-excitable cell types and are found in organisms ranging from Hydra vulgaris to Homo sapiens. Although they are firmly in the superfamily of S4 domain-bearing voltage-sensing ion channels, KCNQ channels are highly sensitive to a range of endogenous and exogenous small molecules that act directly on the pore, the voltage-sensing domain, or the interface between the two. The focus of this review is regulation of KCNQs by direct binding of neurotransmitters and metabolites from both animals and plants and the role of the latter in the effects of plants consumed for food and as traditional folk medicines. The conceptual question arises: Are KCNQs voltage-gated channels that are also sensitive to ligands or ligand-gated channels that are also sensitive to voltage?
Keywords: GABA; KCNE; KCNQ2; KCNQ3; KCNQ5; epilepsy; herbal medicine; hypertension.
Copyright © 2020 Abbott.
Figures





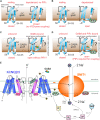
References
-
- Abbott G. W. (2014). Biology of the KCNQ1 potassium channel. New J. Sci. 2014:237431.
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

