Personalised mapping of tumour development in synchronous colorectal cancer patients
- PMID: 32655884
- PMCID: PMC7335056
- DOI: 10.1038/s41525-020-0134-3
Personalised mapping of tumour development in synchronous colorectal cancer patients
Abstract
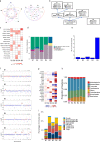
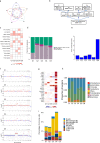
Synchronous colorectal cancers (syCRCs) are two or more primary tumours identified simultaneously in a patient. Previous studies report high inter-tumour heterogeneity between syCRCs, suggesting independent origin and different treatment response, making their management particularly challenging, with no specific guidelines currently in place. Here, we performed in-depth bioinformatic analyses of genomic and transcriptomic data of a total of eleven syCRCs and one metachronous CRC collected from three patients. We found mixed microsatellite status between and within patients. Overlap of mutations between synchronous tumours was consistently low (<0.5%) and heterogeneity of driver events across syCRCs was high in all patients. Microbial analysis revealed the presence of Fusobacterium nucleatum species in patients with MSI tumours, while quantification of tumour immune infiltration showed varying immune responses between syCRCs. Our results suggest high heterogeneity of syCRCs within patients but find clinically actionable biomarkers that help predict responses to currently available targeted therapies. Our study highlights the importance of personalised genome and transcriptome sequencing of all synchronous lesions to aid therapy decision and improve management of syCRC patients.
Keywords: Cancer genomics; Colorectal cancer; Genome informatics.
© The Author(s) 2020.
Conflict of interest statement
Competing interestsThe authors declare no competing interests.
Figures

References
Publication types
LinkOut - more resources
Full Text Sources

