Deep immune profiling of COVID-19 patients reveals distinct immunotypes with therapeutic implications
- PMID: 32669297
- PMCID: PMC7402624
- DOI: 10.1126/science.abc8511
Deep immune profiling of COVID-19 patients reveals distinct immunotypes with therapeutic implications
Abstract
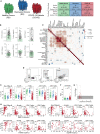
Coronavirus disease 2019 (COVID-19) is currently a global pandemic, but human immune responses to the virus remain poorly understood. We used high-dimensional cytometry to analyze 125 COVID-19 patients and compare them with recovered and healthy individuals. Integrated analysis of ~200 immune and ~50 clinical features revealed activation of T cell and B cell subsets in a proportion of patients. A subgroup of patients had T cell activation characteristic of acute viral infection and plasmablast responses reaching >30% of circulating B cells. However, another subgroup had lymphocyte activation comparable with that in uninfected individuals. Stable versus dynamic immunological signatures were identified and linked to trajectories of disease severity change. Our analyses identified three immunotypes associated with poor clinical trajectories versus improving health. These immunotypes may have implications for the design of therapeutics and vaccines for COVID-19.
Copyright © 2020 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Figures






Update of
-
Deep immune profiling of COVID-19 patients reveals patient heterogeneity and distinct immunotypes with implications for therapeutic interventions.bioRxiv [Preprint]. 2020 May 23:2020.05.20.106401. doi: 10.1101/2020.05.20.106401. bioRxiv. 2020. Update in: Science. 2020 Sep 4;369(6508):eabc8511. doi: 10.1126/science.abc8511. PMID: 32511371 Free PMC article. Updated. Preprint.
References
-
- E. Iype, S. Gulati, Understanding the asymmetric spread and case fatality rate (CFR) for COVID-19 among countries. medRxiv 20073791 [Preprint]. 26 April 2020. 10.1101/2020.04.21.20073791 - DOI
-
- Vabret N., Britton G. J., Gruber C., Hegde S., Kim J., Kuksin M., Levantovsky R., Malle L., Moreira A., Park M. D., Pia L., Risson E., Saffern M., Salomé B., Esai Selvan M., Spindler M. P., Tan J., van der Heide V., Gregory J. K., Alexandropoulos K., Bhardwaj N., Brown B. D., Greenbaum B., Gümüş Z. H., Homann D., Horowitz A., Kamphorst A. O., Curotto de Lafaille M. A., Mehandru S., Merad M., Samstein R. M.; Sinai Immunology Review Project , Immunology of COVID-19: Current state of the science. Immunity 52, 910–941 (2020). 10.1016/j.immuni.2020.05.002 - DOI - PMC - PubMed
-
- D. Weiskopf, K. S. Schmitz, M. P. Raadsen, A. Grifoni, N. M. A. Okba, H. Endeman, J. P. C. van den Akker, R. Molenkamp, M. P. G. Koopmans, E. C. M. van Gorp, B. L. Haagmans, R. L. de Swart, A. Sette, R. D. de Vries, Phenotype of SARS-CoV-2-specific T-cells in COVID-19 patients with acute respiratory distress syndrome. medRxiv 20062349 [Preprint]. 29 May 2020. 10.1101/2020.04.11.20062349 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

