G4C2 Repeat RNA Initiates a POM121-Mediated Reduction in Specific Nucleoporins in C9orf72 ALS/FTD
- PMID: 32673563
- PMCID: PMC8077944
- DOI: 10.1016/j.neuron.2020.06.027
G4C2 Repeat RNA Initiates a POM121-Mediated Reduction in Specific Nucleoporins in C9orf72 ALS/FTD
Abstract
Through mechanisms that remain poorly defined, defects in nucleocytoplasmic transport and accumulations of specific nuclear-pore-complex-associated proteins have been reported in multiple neurodegenerative diseases, including C9orf72 Amyotrophic Lateral Sclerosis and Frontotemporal Dementia (ALS/FTD). Using super-resolution structured illumination microscopy, we have explored the mechanism by which nucleoporins are altered in nuclei isolated from C9orf72 induced pluripotent stem-cell-derived neurons (iPSNs). Of the 23 nucleoporins evaluated, we observed a reduction in a subset of 8, including key components of the nuclear pore complex scaffold and the transmembrane nucleoporin POM121. Reduction in POM121 appears to initiate a decrease in the expression of seven additional nucleoporins, ultimately affecting the localization of Ran GTPase and subsequent cellular toxicity in C9orf72 iPSNs. Collectively, our data suggest that the expression of expanded C9orf72 ALS/FTD repeat RNA alone affects nuclear POM121 expression in the initiation of a pathological cascade affecting nucleoporin levels within neuronal nuclei and ultimately downstream neuronal survival.
Keywords: ALS; C9orf72; FTD; POM121; neurodegeneration; nuclear pore complex.
Copyright © 2020 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declarations of Interests The authors declare no competing interests.
Figures
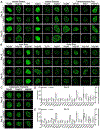






References
-
- Ababneh NA, Scaber J, Flynn R, Douglas A, Barbagallo P, Candalija A, Turner MR, Sims D, Dafinca R, Cowley SA, et al. (2020). Correction of amyotrophic lateral sclerosis related phenotypes in induced pluripotent stem cell-derived motor neurons carrying a hexanucleotide expansion mutation in C9orf72 by CRISPR/Cas9 genome editing using homology-directed repair. Human molecular genetics. - PMC - PubMed
-
- Antonin W, Franz C, Haselmann U, Antony C, and Mattaj IW (2005). The integral membrane nucleoporin pom121 functionally links nuclear pore complex assembly and nuclear envelope formation. Molecular cell 17, 83–92. - PubMed
-
- Beck M, and Hurt E (2017). The nuclear pore complex: understanding its function through structural insight. Nature reviews Molecular cell biology 18, 73–89. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

