This is a preprint.
Comprehensive in-vivo secondary structure of the SARS-CoV-2 genome reveals novel regulatory motifs and mechanisms
- PMID: 32676598
- PMCID: PMC7359520
- DOI: 10.1101/2020.07.10.197079
Comprehensive in-vivo secondary structure of the SARS-CoV-2 genome reveals novel regulatory motifs and mechanisms
Update in
-
Comprehensive in vivo secondary structure of the SARS-CoV-2 genome reveals novel regulatory motifs and mechanisms.Mol Cell. 2021 Feb 4;81(3):584-598.e5. doi: 10.1016/j.molcel.2020.12.041. Epub 2021 Jan 1. Mol Cell. 2021. PMID: 33444546 Free PMC article.
Abstract
SARS-CoV-2 is the positive-sense RNA virus that causes COVID-19, a disease that has triggered a major human health and economic crisis. The genome of SARS-CoV-2 is unique among viral RNAs in its vast potential to form stable RNA structures and yet, as much as 97% of its 30 kilobases have not been structurally explored in the context of a viral infection. Our limited knowledge of SARS-CoV-2 genomic architecture is a fundamental limitation to both our mechanistic understanding of coronavirus life cycle and the development of COVID-19 RNA-based therapeutics. Here, we apply a novel long amplicon strategy to determine for the first time the secondary structure of the SARS-CoV-2 RNA genome probed in infected cells. In addition to the conserved structural motifs at the viral termini, we report new structural features like a conformationally flexible programmed ribosomal frameshifting pseudoknot, and a host of novel RNA structures, each of which highlights the importance of studying viral structures in their native genomic context. Our in-depth structural analysis reveals extensive networks of well-folded RNA structures throughout Orf1ab and reveals new aspects of SARS-CoV-2 genome architecture that distinguish it from other single-stranded, positive-sense RNA viruses. Evolutionary analysis of RNA structures in SARS-CoV-2 shows that several features of its genomic structure are conserved across beta coronaviruses and we pinpoint individual regions of well-folded RNA structure that merit downstream functional analysis. The native, complete secondary structure of SAR-CoV-2 presented here is a roadmap that will facilitate focused studies on mechanisms of replication, translation and packaging, and guide the identification of new RNA drug targets against COVID-19.
Conflict of interest statement
Declaration of Interests
A patent application on MarathonRT has been filed by Yale University.
Figures




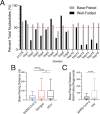


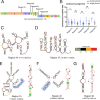
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
