Distinct and stage-specific contributions of TET1 and TET2 to stepwise cytosine oxidation in the transition from naive to primed pluripotency
- PMID: 32694513
- PMCID: PMC7374584
- DOI: 10.1038/s41598-020-68600-3
Distinct and stage-specific contributions of TET1 and TET2 to stepwise cytosine oxidation in the transition from naive to primed pluripotency
Abstract
Cytosine DNA bases can be methylated by DNA methyltransferases and subsequently oxidized by TET proteins. The resulting 5-hydroxymethylcytosine (5hmC), 5-formylcytosine (5fC), and 5-carboxylcytosine (5caC) are considered demethylation intermediates as well as stable epigenetic marks. To dissect the contributions of these cytosine modifying enzymes, we generated combinations of Tet knockout (KO) embryonic stem cells (ESCs) and systematically measured protein and DNA modification levels at the transition from naive to primed pluripotency. Whereas the increase of genomic 5-methylcytosine (5mC) levels during exit from pluripotency correlated with an upregulation of the de novo DNA methyltransferases DNMT3A and DNMT3B, the subsequent oxidation steps turned out to be far more complex. The strong increase of oxidized cytosine bases (5hmC, 5fC, and 5caC) was accompanied by a drop in TET2 levels, yet the analysis of KO cells suggested that TET2 is responsible for most 5fC formation. The comparison of modified cytosine and enzyme levels in Tet KO cells revealed distinct and differentiation-dependent contributions of TET1 and TET2 to 5hmC and 5fC formation arguing against a processive mechanism of 5mC oxidation. The apparent independent steps of 5hmC and 5fC formation suggest yet to be identified mechanisms regulating TET activity that may constitute another layer of epigenetic regulation.
Conflict of interest statement
The authors declare no competing interests.
Figures
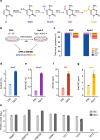


References
-
- Smith ZD, Meissner A. DNA methylation: Roles in mammalian development. Nat. Rev. Genet. 2013;14:204–220. - PubMed
-
- Monk M, Boubelik M, Lehnert S. Temporal and regional changes in DNA methylation in the embryonic, extraembryonic and germ cell lineages during mouse embryo development. Development. 1987;99:371–382. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

