Mycobacterium tuberculosis FasR senses long fatty acyl-CoA through a tunnel and a hydrophobic transmission spine
- PMID: 32710080
- PMCID: PMC7382501
- DOI: 10.1038/s41467-020-17504-x
Mycobacterium tuberculosis FasR senses long fatty acyl-CoA through a tunnel and a hydrophobic transmission spine
Abstract
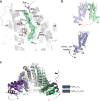
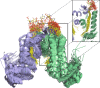
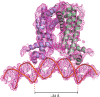
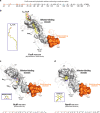
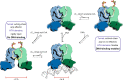
Mycobacterium tuberculosis is a pathogen with a unique cell envelope including very long fatty acids, implicated in bacterial resistance and host immune modulation. FasR is a TetR-like transcriptional activator that plays a central role in sensing mycobacterial long-chain fatty acids and regulating lipid biosynthesis. Here we disclose crystal structures of M. tuberculosis FasR in complex with acyl effector ligands and with DNA, uncovering its molecular sensory and switching mechanisms. A long tunnel traverses the entire effector-binding domain, enabling long fatty acyl effectors to bind. Only when the tunnel is entirely occupied, the protein dimer adopts a rigid configuration with its DNA-binding domains in an open state, leading to DNA dissociation. The protein-folding hydrophobic core connects the two domains, and is completed into a continuous spine when the effector binds. Such a transmission spine is conserved in a large number of TetR-like regulators, offering insight into effector-triggered allosteric functional control.
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Crystal structure of FadR, a fatty acid-responsive transcription factor with a novel acyl coenzyme A-binding fold.EMBO J. 2000 Oct 2;19(19):5167-77. doi: 10.1093/emboj/19.19.5167. EMBO J. 2000. PMID: 11013219 Free PMC article.
-
MadR mediates acyl CoA-dependent regulation of mycolic acid desaturation in mycobacteria.Proc Natl Acad Sci U S A. 2022 Feb 22;119(8):e2111059119. doi: 10.1073/pnas.2111059119. Proc Natl Acad Sci U S A. 2022. PMID: 35165190 Free PMC article.
-
Identification and characterization of Rv0494: a fatty acid-responsive protein of the GntR/FadR family from Mycobacterium tuberculosis.Microbiology (Reading). 2013 May;159(Pt 5):913-923. doi: 10.1099/mic.0.066654-0. Epub 2013 Mar 8. Microbiology (Reading). 2013. PMID: 23475950
-
Long-chain acyl-CoA-dependent regulation of gene expression in bacteria, yeast and mammals.J Nutr. 2000 Feb;130(2S Suppl):305S-309S. doi: 10.1093/jn/130.2.305S. J Nutr. 2000. PMID: 10721893 Review.
-
Fatty acyl-AMP ligases and polyketide synthases are unique enzymes of lipid biosynthetic machinery in Mycobacterium tuberculosis.Tuberculosis (Edinb). 2011 Sep;91(5):448-55. doi: 10.1016/j.tube.2011.04.006. Epub 2011 May 20. Tuberculosis (Edinb). 2011. PMID: 21601529 Review.
Cited by
-
Pseudomonas aeruginosa regulator PvrA binds simultaneously to multiple pseudo-palindromic sites for efficient transcription activation.Sci China Life Sci. 2024 May;67(5):900-912. doi: 10.1007/s11427-022-2363-y. Epub 2023 Sep 26. Sci China Life Sci. 2024. PMID: 37938507
-
Structures of the DarR transcription regulator reveal unique modes of second messenger and DNA binding.Nat Commun. 2023 Nov 9;14(1):7239. doi: 10.1038/s41467-023-42823-0. Nat Commun. 2023. PMID: 37945601 Free PMC article.
-
The Lack of the TetR-Like Repressor Gene BCG_2177c (Rv2160A) May Help Mycobacteria Overcome Intracellular Redox Stress and Survive Longer Inside Macrophages When Surrounded by a Lipid Environment.Front Cell Infect Microbiol. 2022 Jul 7;12:907890. doi: 10.3389/fcimb.2022.907890. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 35873160 Free PMC article.
-
Genetically encoded transcriptional plasticity underlies stress adaptation in Mycobacterium tuberculosis.bioRxiv [Preprint]. 2023 Aug 20:2023.08.20.553992. doi: 10.1101/2023.08.20.553992. bioRxiv. 2023. Update in: Nat Commun. 2024 Apr 10;15(1):3088. doi: 10.1038/s41467-024-47410-5. PMID: 37645742 Free PMC article. Updated. Preprint.
-
Genetically encoded transcriptional plasticity underlies stress adaptation in Mycobacterium tuberculosis.Res Sq [Preprint]. 2023 Sep 18:rs.3.rs-3303807. doi: 10.21203/rs.3.rs-3303807/v1. Res Sq. 2023. Update in: Nat Commun. 2024 Apr 10;15(1):3088. doi: 10.1038/s41467-024-47410-5. PMID: 37790329 Free PMC article. Updated. Preprint.
References
-
- Daffé, M. & Draper, P. in Advances in Microbial Physiology Vol. 39 (ed. Poole, R. K.) 131–203 (Academic Press, US, 1997).
-
- Bhatt A, Molle V, Besra GS, Jacobs WR, Jr., Kremer L. The Mycobacterium tuberculosis FAS-II condensing enzymes: their role in mycolic acid biosynthesis, acid-fastness, pathogenesis and in future drug development. Mol. Microbiol. 2007;64:1442–1454. - PubMed
-
- Marrakchi H, Laneelle MA, Daffe M. Mycolic acids: structures, biosynthesis, and beyond. Chem. Biol. 2014;21:67–85. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

