Genome-wide R-loop analysis defines unique roles for DDX5, XRN2, and PRMT5 in DNA/RNA hybrid resolution
- PMID: 32747416
- PMCID: PMC7409538
- DOI: 10.26508/lsa.202000762
Genome-wide R-loop analysis defines unique roles for DDX5, XRN2, and PRMT5 in DNA/RNA hybrid resolution
Abstract
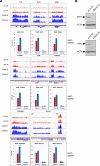
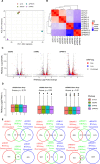
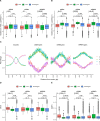
DDX5, XRN2, and PRMT5 have been shown to resolve DNA/RNA hybrids (R-loops) at RNA polymerase II transcription termination sites at few genomic loci. Herein, we perform genome-wide R-loop mapping using classical DNA/RNA immunoprecipitation and high-throughput sequencing (DRIP-seq) of loci regulated by DDX5, XRN2, and PRMT5. We observed hundreds to thousands of R-loop gains and losses at transcribed loci in DDX5-, XRN2-, and PRMT5-deficient U2OS cells. R-loop gains were characteristic of highly transcribed genes located at gene-rich regions, whereas R-loop losses were observed in low-density gene areas. DDX5, XRN2, and PRMT5 shared many R-loop gain loci at transcription termination sites, consistent with their coordinated role in RNA polymerase II transcription termination. DDX5-depleted cells had unique R-loop gain peaks near the transcription start site that did not overlap with those of siXRN2 and siPRMT5 cells, suggesting a role for DDX5 in transcription initiation independent of XRN2 and PRMT5. Moreover, we observed that the accumulated R-loops at certain loci in siDDX5, siXRN2, and siPRMT5 cells near the transcription start site of genes led to antisense intergenic transcription. Our findings define unique and shared roles of DDX5, XRN2, and PRMT5 in DNA/RNA hybrid regulation.
© 2020 Villarreal et al.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures







Similar articles
-
Arginine methylation of the DDX5 helicase RGG/RG motif by PRMT5 regulates resolution of RNA:DNA hybrids.EMBO J. 2019 Aug 1;38(15):e100986. doi: 10.15252/embj.2018100986. Epub 2019 Jun 21. EMBO J. 2019. PMID: 31267554 Free PMC article.
-
DDX21 mediates co-transcriptional RNA m6A modification to promote transcription termination and genome stability.Mol Cell. 2024 May 2;84(9):1711-1726.e11. doi: 10.1016/j.molcel.2024.03.006. Epub 2024 Apr 2. Mol Cell. 2024. PMID: 38569554
-
Thrap3 promotes R-loop resolution via interaction with methylated DDX5.Exp Mol Med. 2021 Oct;53(10):1602-1611. doi: 10.1038/s12276-021-00689-6. Epub 2021 Oct 25. Exp Mol Med. 2021. PMID: 34697388 Free PMC article.
-
A genome-wide and cotranscriptional suppressor of R loops.Genes Dev. 2020 Jul 1;34(13-14):863-864. doi: 10.1101/gad.339861.120. Genes Dev. 2020. PMID: 32611612 Free PMC article. Review.
-
Regulatory R-loops as facilitators of gene expression and genome stability.Nat Rev Mol Cell Biol. 2020 Mar;21(3):167-178. doi: 10.1038/s41580-019-0206-3. Epub 2020 Jan 31. Nat Rev Mol Cell Biol. 2020. PMID: 32005969 Free PMC article. Review.
Cited by
-
High PRMT5 levels, maintained by KEAP1 inhibition, drive chemoresistance in high-grade serous ovarian cancer.J Clin Invest. 2025 Mar 17;135(6):e184283. doi: 10.1172/JCI184283. J Clin Invest. 2025. PMID: 40091834 Free PMC article.
-
Approaches for Mapping and Analysis of R-loops.Curr Protoc. 2024 Apr;4(4):e1037. doi: 10.1002/cpz1.1037. Curr Protoc. 2024. PMID: 38666626 Free PMC article. Review.
-
Molecular Basis of XRN2-Deficient Cancer Cell Sensitivity to Poly(ADP-ribose) Polymerase Inhibition.Cancers (Basel). 2024 Jan 30;16(3):595. doi: 10.3390/cancers16030595. Cancers (Basel). 2024. PMID: 38339346 Free PMC article.
-
Multifunctional role of DEAD-box helicase 41 in innate immunity, hematopoiesis and disease.Front Immunol. 2024 Aug 9;15:1451705. doi: 10.3389/fimmu.2024.1451705. eCollection 2024. Front Immunol. 2024. PMID: 39185415 Free PMC article. Review.
-
p53 activates circASCC3 to repress R-loops and enhance resistance to chemotherapy.Proc Natl Acad Sci U S A. 2025 Mar 18;122(11):e2415869122. doi: 10.1073/pnas.2415869122. Epub 2025 Mar 11. Proc Natl Acad Sci U S A. 2025. PMID: 40067902 Free PMC article.
References
-
- Aronesty E. (2013) Comparison of sequencing utility programs. Open Bioinforma J 7: 1–8. 10.2174/1875036201307010001 - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
