On the evolutionary epidemiology of SARS-CoV-2
- PMID: 32750338
- PMCID: PMC7287426
- DOI: 10.1016/j.cub.2020.06.031
On the evolutionary epidemiology of SARS-CoV-2
Abstract
There is no doubt that the novel coronavirus SARS-CoV-2 that causes COVID-19 is mutating and thus has the potential to adapt during the current pandemic. Whether this evolution will lead to changes in the transmission, the duration, or the severity of the disease is not clear. This has led to considerable scientific and media debate, from raising alarms about evolutionary change to dismissing it. Here we review what little is currently known about the evolution of SARS-CoV-2 and extend existing evolutionary theory to consider how selection might be acting upon the virus during the COVID-19 pandemic. Although there is currently no definitive evidence that SARS-CoV-2 is undergoing further adaptation, continued evidence-based analysis of evolutionary change is important so that public health measures can be adjusted in response to substantive changes in the infectivity or severity of COVID-19.
Copyright © 2020 Elsevier Inc. All rights reserved.
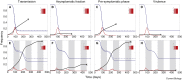
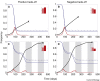
Figures


References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

