Phage-Antibiotic Synergy Is Driven by a Unique Combination of Antibacterial Mechanism of Action and Stoichiometry
- PMID: 32753497
- PMCID: PMC7407087
- DOI: 10.1128/mBio.01462-20
Phage-Antibiotic Synergy Is Driven by a Unique Combination of Antibacterial Mechanism of Action and Stoichiometry
Abstract
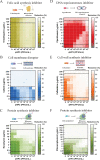
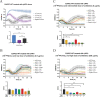
The continued rise in antibiotic resistance is precipitating a medical crisis. Bacteriophage (phage) has been hailed as one possible therapeutic option to augment the efficacy of antibiotics. However, only a few studies have addressed the synergistic relationship between phage and antibiotics. Here, we report a comprehensive analysis of phage-antibiotic interaction that evaluates synergism, additivism, and antagonism for all classes of antibiotics across clinically achievable stoichiometries. We combined an optically based real-time microtiter plate readout with a matrix-like heat map of treatment potencies to measure phage and antibiotic synergy (PAS), a process we term synography. Phage-antibiotic synography was performed against a pandemic drug-resistant clonal group of extraintestinal pathogenic Escherichia coli (ExPEC) with antibiotic levels blanketing the MIC across seven orders of viral titers. Our results suggest that, under certain conditions, phages provide an adjuvating effect by lowering the MIC for drug-resistant strains. Furthermore, synergistic and antagonistic interactions are highly dependent on the mechanism of bacterial inhibition by the class of antibiotic paired to the phage, and when synergism is observed, it suppresses the emergence of resistant cells. Host conditions that simulate the infection environment, including serum and urine, suppress PAS in a bacterial growth-dependent manner. Lastly, two different related phages that differed in their burst sizes produced drastically different synograms. Collectively, these data suggest lytic phages can resuscitate an ineffective antibiotic for previously resistant bacteria while also synergizing with antibiotics in a class-dependent manner, processes that may be dampened by lower bacterial growth rates found in host environments.IMPORTANCE Bacteriophage (phage) therapy is a promising approach to combat the rise of multidrug-resistant bacteria. Currently, the preferred clinical modality is to pair phage with an antibiotic, a practice thought to improve efficacy. However, antagonism between phage and antibiotics has been reported, the choice of phage and antibiotic is not often empirically determined, and the effect of the host factors on the effectiveness is unknown. Here, we interrogate phage-antibiotic interactions across antibiotics with different mechanisms of action. Our results suggest that phage can lower the working MIC for bacterial strains already resistant to the antibiotic, is dependent on the antibiotic class and stoichiometry of the pairing, and is dramatically influenced by the host microenvironment.
Keywords: Escherichia coli; adjuvant; antibiotic; bacteriophage; clinical isolate; combinatorial treatment; phage; phage therapy; synergy; synogram; synography.
Copyright © 2020 Gu Liu et al.
Figures




References
-
- Centers for Disease Control and Prevention. 2019. Antibiotic resistance threats in the United States, 2019. Centers for Disease Control and Prevention, Atlanta, GA.
-
- World Health Organization. 2014. Antimicrobial resistance: global report on surveillance. World Health Organization, Geneva, Switzerland.
-
- O’Neill J. 2016. Tackling drug-resistant infections globally: final report and recommendations. https://amr-review.org/sites/default/files/160525_Final%20paper_with%20c....
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

