The First Nonmammalian Pegivirus Demonstrates Efficient In Vitro Replication and High Lymphotropism
- PMID: 32759314
- PMCID: PMC7527067
- DOI: 10.1128/JVI.01150-20
The First Nonmammalian Pegivirus Demonstrates Efficient In Vitro Replication and High Lymphotropism
Abstract
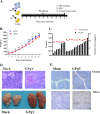
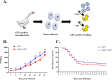
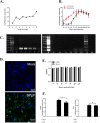
Members of the Pegivirus genus, family Flaviviridae, widely infect humans and other mammals, including nonhuman primates, bats, horses, pigs, and rodents, but are not associated with disease. Here, we report a new, genetically distinct pegivirus in goose (Anser cygnoides), the first identified in a nonmammalian host species. Goose pegivirus (GPgV) can be propagated in goslings, embryonated goose eggs, and primary goose embryo fibroblasts, and is thus the first pegivirus that can be efficiently cultured in vitro Experimental infection of GPgV in goslings via intravenous injection revealed robust replication and high lymphotropism. Analysis of the tissue tropism of GPgV revealed that the spleen and thymus were the organs bearing the highest viral loads. Importantly, GPgV could promote clinical manifestations of goose parvovirus infection, including reduced weight gain and 7% mortality. This finding contrasts with the lack of pathogenicity that is characteristic of previously reported pegiviruses.IMPORTANCE Members of the Pegivirus genus, family Flaviviridae, widely infect humans and other mammals, but are described as causing persistent infection and lacking pathogenicity. The efficiency of in vitro replication systems for pegivirus is poor, thus limiting investigation into viral replication steps. Because of that, the pathogenesis, cellular tropism, route of transmission, biology, and epidemiology of pegiviruses remain largely uncovered. Here, we report a phylogenetically distinct goose pegivirus (GPgV) that should be classified as a new species. GPgV proliferated in cell culture in a species- and cell-type-specific manner. Animal experiments show GPgV lymphotropism and promote goose parvovirus clinical manifestations. This study provides the first cell culture model for pegivirus, opening new possibilities for studies of pegivirus molecular biology. More importantly, our findings stand in contrast to the lack of identified pathogenicity of previously reported pegiviruses, which sheds lights on the pathobiology of pegivirus.
Keywords: goose; pegivirus.
Copyright © 2020 American Society for Microbiology.
Figures




References
-
- Kapoor A, Simmonds P, Scheel TK, Hjelle B, Cullen JM, Burbelo PD, Chauhan LV, Duraisamy R, Sanchez Leon M, Jain K, Vandegrift KJ, Calisher CH, Rice CM, Lipkin WI. 2013. Identification of rodent homologs of hepatitis C virus and pegiviruses. mBio 4:e00216-13–e00213. doi:10.1128/mBio.00216-13. - DOI - PMC - PubMed
-
- Smith DB, Becher P, Bukh J, Gould EA, Meyers G, Monath T, Muerhoff AS, Pletnev A, Rico-Hesse R, Stapleton JT, Simmonds P. 2016. Proposed update to the taxonomy of the genera Hepacivirus and Pegivirus within the Flaviviridae family. J Gen Virol 97:2894–2907. doi:10.1099/jgv.0.000612. - DOI - PMC - PubMed
-
- Chandriani S, Skewes-Cox P, Zhong W, Ganem DE, Divers TJ, Van Blaricum AJ, Tennant BC, Kistler AL. 2013. Identification of a previously undescribed divergent virus from the Flaviviridae family in an outbreak of equine serum hepatitis. Proc Natl Acad Sci U S A 110:E1407–E1415. doi:10.1073/pnas.1219217110. - DOI - PMC - PubMed
-
- Bailey AL, Buechler CR, Matson DR, Peterson EJ, Brunner KG, Mohns MS, Breitbach M, Stewart LM, Ericsen AJ, Newman CM, Koenig MR, Mohr E, Tan J, Capuano S, Simmons HA, Yang DT, O'Connor DH. 2017. Pegivirus avoids immune recognition but does not attenuate acute-phase disease in a macaque model of HIV infection. PLoS Pathog 13:e1006692. doi:10.1371/journal.ppat.1006692. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

