The histone methyltransferase DOT1L prevents antigen-independent differentiation and safeguards epigenetic identity of CD8+ T cells
- PMID: 32764145
- PMCID: PMC7456197
- DOI: 10.1073/pnas.1920372117
The histone methyltransferase DOT1L prevents antigen-independent differentiation and safeguards epigenetic identity of CD8+ T cells
Abstract
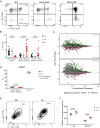
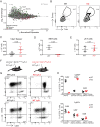
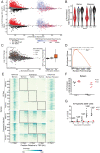
Cytotoxic T cell differentiation is guided by epigenome adaptations, but how epigenetic mechanisms control lymphocyte development has not been well defined. Here we show that the histone methyltransferase DOT1L, which marks the nucleosome core on active genes, safeguards normal differentiation of CD8+ T cells. T cell-specific ablation of Dot1L resulted in loss of naïve CD8+ T cells and premature differentiation toward a memory-like state, independent of antigen exposure and in a cell-intrinsic manner. Mechanistically, DOT1L controlled CD8+ T cell differentiation by ensuring normal T cell receptor density and signaling. DOT1L also maintained epigenetic identity, in part by indirectly supporting the repression of developmentally regulated genes. Finally, deletion of Dot1L in T cells resulted in an impaired immune response. Through our study, DOT1L is emerging as a central player in physiology of CD8+ T cells, acting as a barrier to prevent premature differentiation and controlling epigenetic integrity.
Keywords: DOT1L; H3K79me2; T cell; epigenetics; virtual memory.
Copyright © 2020 the Author(s). Published by PNAS.
Conflict of interest statement
The authors declare no competing interest.
Figures

References
-
- Borst J., Jacobs H., Brouns G., Composition and function of T-cell receptor and B-cell receptor complexes on precursor lymphocytes. Curr. Opin. Immunol. 8, 181–190 (1996). - PubMed
-
- Hussain T., Quinn K. M., Similar but different: Virtual memory CD8 T cells as a memory-like cell population. Immunol. Cell Biol. 97, 675–684 (2019). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

