Differentiating Drosophila female germ cells initiate Polycomb silencing by regulating PRC2-interacting proteins
- PMID: 32773039
- PMCID: PMC7438113
- DOI: 10.7554/eLife.56922
Differentiating Drosophila female germ cells initiate Polycomb silencing by regulating PRC2-interacting proteins
Abstract
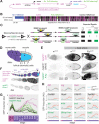
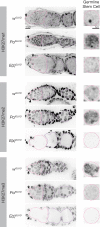
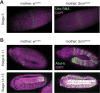
Polycomb silencing represses gene expression and provides a molecular memory of chromatin state that is essential for animal development. We show that Drosophila female germline stem cells (GSCs) provide a powerful system for studying Polycomb silencing. GSCs have a non-canonical distribution of PRC2 activity and lack silenced chromatin like embryonic progenitors. As GSC daughters differentiate into nurse cells and oocytes, nurse cells, like embryonic somatic cells, silence genes in traditional Polycomb domains and in generally inactive chromatin. Developmentally controlled expression of two Polycomb repressive complex 2 (PRC2)-interacting proteins, Pcl and Scm, initiate silencing during differentiation. In GSCs, abundant Pcl inhibits PRC2-dependent silencing globally, while in nurse cells Pcl declines and newly induced Scm concentrates PRC2 activity on traditional Polycomb domains. Our results suggest that PRC2-dependent silencing is developmentally regulated by accessory proteins that either increase the concentration of PRC2 at target sites or inhibit the rate that PRC2 samples chromatin.
Keywords: D. melanogaster; PRC2; chromatin; chromosomes; differentiation; epigenetic; gene expression; nurse cell; polycomb.
© 2020, DeLuca et al.
Conflict of interest statement
SD, MG, LP, AS No competing interests declared
Figures







Similar articles
-
Interdependence of PRC1 and PRC2 for recruitment to Polycomb Response Elements.Nucleic Acids Res. 2016 Dec 1;44(21):10132-10149. doi: 10.1093/nar/gkw701. Epub 2016 Aug 23. Nucleic Acids Res. 2016. PMID: 27557709 Free PMC article.
-
Pcl-PRC2 is needed to generate high levels of H3-K27 trimethylation at Polycomb target genes.EMBO J. 2007 Sep 19;26(18):4078-88. doi: 10.1038/sj.emboj.7601837. Epub 2007 Aug 30. EMBO J. 2007. PMID: 17762866 Free PMC article.
-
Global changes of H3K27me3 domains and Polycomb group protein distribution in the absence of recruiters Spps or Pho.Proc Natl Acad Sci U S A. 2018 Feb 20;115(8):E1839-E1848. doi: 10.1073/pnas.1716299115. Epub 2018 Feb 5. Proc Natl Acad Sci U S A. 2018. PMID: 29432187 Free PMC article.
-
Recruiting polycomb to chromatin.Int J Biochem Cell Biol. 2015 Oct;67:177-87. doi: 10.1016/j.biocel.2015.05.006. Epub 2015 May 14. Int J Biochem Cell Biol. 2015. PMID: 25982201 Free PMC article. Review.
-
Tissue-Specific Tumour Suppressor and Oncogenic Activities of the Polycomb-like Protein MTF2.Genes (Basel). 2023 Sep 27;14(10):1879. doi: 10.3390/genes14101879. Genes (Basel). 2023. PMID: 37895228 Free PMC article. Review.
Cited by
-
Genetic variation in P-element dysgenic sterility is associated with double-strand break repair and alternative splicing of TE transcripts.PLoS Genet. 2022 Dec 7;18(12):e1010080. doi: 10.1371/journal.pgen.1010080. eCollection 2022 Dec. PLoS Genet. 2022. PMID: 36477699 Free PMC article.
-
Chromatin and gene expression changes during female Drosophila germline stem cell development illuminate the biology of highly potent stem cells.Elife. 2023 Oct 13;12:RP90509. doi: 10.7554/eLife.90509. Elife. 2023. PMID: 37831064 Free PMC article.
-
Two distinct waves of transcriptome and translatome changes drive Drosophila germline stem cell differentiation.EMBO J. 2024 Apr;43(8):1591-1617. doi: 10.1038/s44318-024-00070-z. Epub 2024 Mar 13. EMBO J. 2024. PMID: 38480936 Free PMC article.
-
Forecasting histone methylation by Polycomb complexes with minute-scale precision.Sci Adv. 2023 Dec 22;9(51):eadj8198. doi: 10.1126/sciadv.adj8198. Epub 2023 Dec 22. Sci Adv. 2023. PMID: 38134278 Free PMC article.
-
Drosophila COMPASS Complex Subunits Set1 and Ash2 Are Required for Oocyte Determination and Maintenance of the Synaptonemal Complex.J Dev Biol. 2025 Aug 19;13(3):30. doi: 10.3390/jdb13030030. J Dev Biol. 2025. PMID: 40843898 Free PMC article.
References
-
- Babenko VN, Makunin IV, Brusentsova IV, Belyaeva ES, Maksimov DA, Belyakin SN, Maroy P, Vasil'eva LA, Zhimulev IF. Paucity and preferential suppression of transgenes in late replication domains of the D. Melanogaster genome. BMC Genomics. 2010;11:318. doi: 10.1186/1471-2164-11-318. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

