Loss of histone H4 lysine 20 trimethylation in osteosarcoma is associated with aberrant expression ofhistone methyltransferase SUV420H2
- PMID: 32774499
- PMCID: PMC7406877
- DOI: 10.3892/ol.2020.11887
Loss of histone H4 lysine 20 trimethylation in osteosarcoma is associated with aberrant expression ofhistone methyltransferase SUV420H2
Abstract
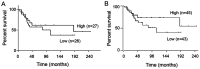
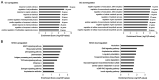
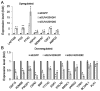
Epigenetic modifications of histones have crucial roles in various types of cancers. The aberrant trimethylation of histone H4 at lysine 20 (H4K20) has been implicated in carcinogenesis. At present, the status of trimethylation at H4k20 (H4K20me3) in osteosarcoma (OS), the predominant bone cancer in humans, is unknown. In the present study, a genome-wide decrease was observed in H4K20me3 levels in OS tissues and cell lines. Reduced levels of lysine methyltransferase 5C (SUV420H2), the histone methyltranferase responsible for modification of H4K20me3, was also observed in OS cells with the associated loss of H4K20me3. Furthermore, a total of 507 SUV420H2-regulated genes were identified through RNA-seq and a number of candidate genes were further validated. Bioinformatic analysis revealed an association between SUV420H2 and multiple signaling pathway, including the mitogen-activated protein kinase, P53, transforming growth factor and the ErbB pathways. These results demonstrated that there are aberrant levels of H4K20me3 and SUV420H2 in OS, and highlighted H4K20me3 as a candidate biomarker for the early detection of OS.
Keywords: epigenetics; histone H4 lysine 20; histone methyltransferase; lysine methyltransferase 5C; osteosarcoma.
Copyright: © Piao et al.
Figures

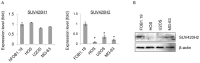

References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
