Extracellular Vesicle and Particle Biomarkers Define Multiple Human Cancers
- PMID: 32795414
- PMCID: PMC7522766
- DOI: 10.1016/j.cell.2020.07.009
Extracellular Vesicle and Particle Biomarkers Define Multiple Human Cancers
Abstract
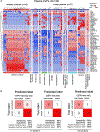
There is an unmet clinical need for improved tissue and liquid biopsy tools for cancer detection. We investigated the proteomic profile of extracellular vesicles and particles (EVPs) in 426 human samples from tissue explants (TEs), plasma, and other bodily fluids. Among traditional exosome markers, CD9, HSPA8, ALIX, and HSP90AB1 represent pan-EVP markers, while ACTB, MSN, and RAP1B are novel pan-EVP markers. To confirm that EVPs are ideal diagnostic tools, we analyzed proteomes of TE- (n = 151) and plasma-derived (n = 120) EVPs. Comparison of TE EVPs identified proteins (e.g., VCAN, TNC, and THBS2) that distinguish tumors from normal tissues with 90% sensitivity/94% specificity. Machine-learning classification of plasma-derived EVP cargo, including immunoglobulins, revealed 95% sensitivity/90% specificity in detecting cancer. Finally, we defined a panel of tumor-type-specific EVP proteins in TEs and plasma, which can classify tumors of unknown primary origin. Thus, EVP proteins can serve as reliable biomarkers for cancer detection and determining cancer type.
Keywords: biomarkers; cancer; cancer of unknown primary origin; damage-associated molecular patterns; early cancer detection; exomeres; exosomes; extracellular vesicles and particles; liquid biopsy; proteomics.
Copyright © 2020 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Interests D.L., A.H., H.S.K., and L.B. have filed a U.S. patent application related to this work.
Figures






Comment in
-
Exosome Profiling Pinpoints Cancer Type.Cancer Discov. 2020 Nov;10(11):1619. doi: 10.1158/2159-8290.CD-NB2020-083. Epub 2020 Sep 4. Cancer Discov. 2020. PMID: 32887700
-
Validation of plasma-derived small extracellular vesicles as cancer biomarkers.Nat Rev Clin Oncol. 2020 Dec;17(12):719-720. doi: 10.1038/s41571-020-00433-5. Nat Rev Clin Oncol. 2020. PMID: 32943766 No abstract available.
-
Extracellular vesicle fingerprinting: the next generation for cancer diagnosis?Signal Transduct Target Ther. 2020 Nov 5;5(1):263. doi: 10.1038/s41392-020-00385-3. Signal Transduct Target Ther. 2020. PMID: 33154355 Free PMC article. No abstract available.
References
-
- Caby MP, Lankar D, Vincendeau-Scherrer C, Raposo G, and Bonnerot C (2005). Exosomal-like vesicles are present in human blood plasma. Int. Immunol 17, 879–887. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- U54 CA163120/CA/NCI NIH HHS/United States
- U54 CA163117/CA/NCI NIH HHS/United States
- R01 CA218513/CA/NCI NIH HHS/United States
- U19 OH009077/OH/NIOSH CDC HHS/United States
- U19 AI144301/AI/NIAID NIH HHS/United States
- P30 CA008748/CA/NCI NIH HHS/United States
- UL1 TR002384/TR/NCATS NIH HHS/United States
- U01 CA210240/CA/NCI NIH HHS/United States
- R01 CA064786/CA/NCI NIH HHS/United States
- R01 CA207983/CA/NCI NIH HHS/United States
- P50 CA127297/CA/NCI NIH HHS/United States
- R50 CA211462/CA/NCI NIH HHS/United States
- S10 RR027699/RR/NCRR NIH HHS/United States
- R01 CA169416/CA/NCI NIH HHS/United States
- U01 CA169538/CA/NCI NIH HHS/United States
- U01 CA224175/CA/NCI NIH HHS/United States
- R01 CA217169/CA/NCI NIH HHS/United States
- R35 CA232093/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

