A comparison of humans and baboons suggests germline mutation rates do not track cell divisions
- PMID: 32804933
- PMCID: PMC7467331
- DOI: 10.1371/journal.pbio.3000838
A comparison of humans and baboons suggests germline mutation rates do not track cell divisions
Abstract
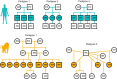
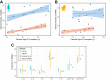
In humans, most germline mutations are inherited from the father. This observation has been widely interpreted as reflecting the replication errors that accrue during spermatogenesis. If so, the male bias in mutation should be substantially lower in a closely related species with similar rates of spermatogonial stem cell divisions but a shorter mean age of reproduction. To test this hypothesis, we resequenced two 3-4 generation nuclear families (totaling 29 individuals) of olive baboons (Papio anubis), who reproduce at approximately 10 years of age on average, and analyzed the data in parallel with three 3-generation human pedigrees (26 individuals). We estimated a mutation rate per generation in baboons of 0.57×10-8 per base pair, approximately half that of humans. Strikingly, however, the degree of male bias in germline mutations is approximately 4:1, similar to that of humans-indeed, a similar male bias is seen across mammals that reproduce months, years, or decades after birth. These results mirror the finding in humans that the male mutation bias is stable with parental ages and cast further doubt on the assumption that germline mutations track cell divisions. Our mutation rate estimates for baboons raise a further puzzle, suggesting a divergence time between apes and Old World monkeys of 65 million years, too old to be consistent with the fossil record; reconciling them now requires not only a slowdown of the mutation rate per generation in humans but also in baboons.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


References
-
- Harland C, Charlier C, Karim L, Cambisano N, Deckers M, Mni M, et al. Frequency of mosaicism points towards mutation-prone early cleavage cell divisions in cattle. bioRxiv [Internet]. 2017;79863 [cited 2018 May 25]. Available from: http://biorxiv.org/content/early/2017/06/29/079863.abstract
-
- Goldberg ME, Harris K. Great ape mutation spectra vary across the phylogeny and the genome due to distinct mutational processes that evolve at different rates. bioRxiv [Internet]. 2019;805598 [cited 2019 Nov 5]. Available from: http://biorxiv.org/content/early/2019/10/15/805598.abstract
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

