The Existence and Localization of Nuclear snoRNAs in Arabidopsis thaliana Revisited
- PMID: 32806552
- PMCID: PMC7464842
- DOI: 10.3390/plants9081016
The Existence and Localization of Nuclear snoRNAs in Arabidopsis thaliana Revisited
Abstract
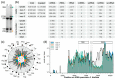
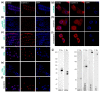
Ribosome biogenesis is one cell function-defining process. It depends on efficient transcription of rDNAs in the nucleolus as well as on the cytosolic synthesis of ribosomal proteins. For newly transcribed rRNA modification and ribosomal protein assembly, so-called small nucleolar RNAs (snoRNAs) and ribosome biogenesis factors (RBFs) are required. For both, an inventory was established for model systems like yeast and humans. For plants, many assignments are based on predictions. Here, RNA deep sequencing after nuclei enrichment was combined with single molecule species detection by northern blot and in vivo fluorescence in situ hybridization (FISH)-based localization studies. In addition, the occurrence and abundance of selected snoRNAs in different tissues were determined. These approaches confirm the presence of most of the database-deposited snoRNAs in cell cultures, but some of them are localized in the cytosol rather than in the nucleus. Further, for the explored snoRNA examples, differences in their abundance in different tissues were observed, suggesting a tissue-specific function of some snoRNAs. Thus, based on prediction and experimental confirmation, many plant snoRNAs can be proposed, while it cannot be excluded that some of the proposed snoRNAs perform alternative functions than are involved in rRNA modification.
Keywords: A. thaliana; NGS; cell fractionation; snoRNAs; tissue specificity.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





Similar articles
-
The Arabidopsis 2'-O-Ribose-Methylation and Pseudouridylation Landscape of rRNA in Comparison to Human and Yeast.Front Plant Sci. 2021 Jul 26;12:684626. doi: 10.3389/fpls.2021.684626. eCollection 2021. Front Plant Sci. 2021. PMID: 34381476 Free PMC article. Review.
-
Proteome distribution between nucleoplasm and nucleolus and its relation to ribosome biogenesis in Arabidopsis thaliana.RNA Biol. 2016;13(4):441-54. doi: 10.1080/15476286.2016.1154252. Epub 2016 Mar 16. RNA Biol. 2016. PMID: 26980300 Free PMC article.
-
Analysis of small nucleolar RNAs reveals unique genetic features in malaria parasites.BMC Genomics. 2009 Feb 7;10:68. doi: 10.1186/1471-2164-10-68. BMC Genomics. 2009. PMID: 19200392 Free PMC article.
-
Arabidopsis small nucleolar RNA monitors the efficient pre-rRNA processing during ribosome biogenesis.Proc Natl Acad Sci U S A. 2016 Oct 18;113(42):11967-11972. doi: 10.1073/pnas.1614852113. Epub 2016 Oct 5. Proc Natl Acad Sci U S A. 2016. PMID: 27708161 Free PMC article.
-
Small nucleolar RNAs: versatile trans-acting molecules of ancient evolutionary origin.Gene Expr. 2002;10(1-2):17-39. Gene Expr. 2002. PMID: 11868985 Free PMC article. Review.
Cited by
-
Mapping rRNA 2'-O-methylations and identification of C/D snoRNAs in Arabidopsis thaliana plants.RNA Biol. 2021 Nov;18(11):1760-1777. doi: 10.1080/15476286.2020.1869892. Epub 2021 Feb 17. RNA Biol. 2021. PMID: 33596769 Free PMC article.
-
EL-RMLocNet: An explainable LSTM network for RNA-associated multi-compartment localization prediction.Comput Struct Biotechnol J. 2022 Jul 26;20:3986-4002. doi: 10.1016/j.csbj.2022.07.031. eCollection 2022. Comput Struct Biotechnol J. 2022. PMID: 35983235 Free PMC article.
-
Low dose ribosomal DNA P-loop mutation affects development and enforces autophagy in Arabidopsis.RNA Biol. 2024 Jan;21(1):1-15. doi: 10.1080/15476286.2023.2298532. Epub 2023 Dec 29. RNA Biol. 2024. PMID: 38156797 Free PMC article.
-
AtSNU13 modulates pre-mRNA splicing of RBOHD and ALD1 to regulate plant immunity.BMC Biol. 2024 Jul 10;22(1):153. doi: 10.1186/s12915-024-01951-9. BMC Biol. 2024. PMID: 38982460 Free PMC article.
-
Cryo-EM structure of wheat ribosome reveals unique features of the plant ribosomes.Structure. 2024 May 2;32(5):562-574.e3. doi: 10.1016/j.str.2024.02.006. Epub 2024 Mar 7. Structure. 2024. PMID: 38458197 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

