Selective inhibition of STAT3 signaling using monobodies targeting the coiled-coil and N-terminal domains
- PMID: 32807795
- PMCID: PMC7431413
- DOI: 10.1038/s41467-020-17920-z
Selective inhibition of STAT3 signaling using monobodies targeting the coiled-coil and N-terminal domains
Abstract
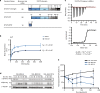
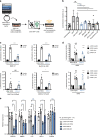
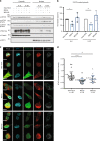
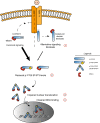
The transcription factor STAT3 is frequently activated in human solid and hematological malignancies and remains a challenging therapeutic target with no approved drugs to date. Here, we develop synthetic antibody mimetics, termed monobodies, to interfere with STAT3 signaling. These monobodies are highly selective for STAT3 and bind with nanomolar affinity to the N-terminal and coiled-coil domains. Interactome analysis detects no significant binding to other STATs or additional off-target proteins, confirming their exquisite specificity. Intracellular expression of monobodies fused to VHL, an E3 ubiquitin ligase substrate receptor, results in degradation of endogenous STAT3. The crystal structure of STAT3 in complex with monobody MS3-6 reveals bending of the coiled-coil domain, resulting in diminished DNA binding and nuclear translocation. MS3-6 expression strongly inhibits STAT3-dependent transcriptional activation and disrupts STAT3 interaction with the IL-22 receptor. Therefore, our study establishes innovative tools to interfere with STAT3 signaling by different molecular mechanisms.
Conflict of interest statement
A.K. and S.K. are listed as inventors on issued and pending patents on the monobody technology filed by The University of Chicago (US Patent 9512199 B2 and related pending applications). The other authors declare no competing interests.
Figures



Similar articles
-
Selective Targeting of SH2 Domain-Phosphotyrosine Interactions of Src Family Tyrosine Kinases with Monobodies.J Mol Biol. 2017 May 5;429(9):1364-1380. doi: 10.1016/j.jmb.2017.03.023. Epub 2017 Mar 25. J Mol Biol. 2017. PMID: 28347651 Free PMC article.
-
Dissection of the BCR-ABL signaling network using highly specific monobody inhibitors to the SHP2 SH2 domains.Proc Natl Acad Sci U S A. 2013 Sep 10;110(37):14924-9. doi: 10.1073/pnas.1303640110. Epub 2013 Aug 26. Proc Natl Acad Sci U S A. 2013. PMID: 23980151 Free PMC article.
-
The intracellular delivery of a recombinant peptide derived from the acidic domain of PIAS3 inhibits STAT3 transactivation and induces tumor cell death.Mol Cancer Res. 2010 Apr;8(4):539-53. doi: 10.1158/1541-7786.MCR-09-0417. Epub 2010 Apr 6. Mol Cancer Res. 2010. PMID: 20371673
-
Monobodies as enabling tools for structural and mechanistic biology.Curr Opin Struct Biol. 2020 Feb;60:167-174. doi: 10.1016/j.sbi.2020.01.015. Epub 2020 Mar 4. Curr Opin Struct Biol. 2020. PMID: 32145686 Free PMC article. Review.
-
Importance of STAT3 signalling in cancer, metastasis and therapeutic interventions.Cell Signal. 2022 Apr;92:110275. doi: 10.1016/j.cellsig.2022.110275. Epub 2022 Feb 3. Cell Signal. 2022. PMID: 35122990 Review.
Cited by
-
Feasibility of the inhibitor development for cancer: A systematic approach for drug design.PLoS One. 2024 Aug 22;19(8):e0306632. doi: 10.1371/journal.pone.0306632. eCollection 2024. PLoS One. 2024. PMID: 39173044 Free PMC article.
-
JAK/STAT: Why choose a classical or an alternative pathway when you can have both?J Cell Mol Med. 2022 Apr;26(7):1865-1875. doi: 10.1111/jcmm.17168. Epub 2022 Mar 3. J Cell Mol Med. 2022. PMID: 35238133 Free PMC article. Review.
-
Joining the PARty: PARP Regulation of KDM5A during DNA Repair (and Transcription?).Bioessays. 2022 Jul;44(7):e2200015. doi: 10.1002/bies.202200015. Epub 2022 May 9. Bioessays. 2022. PMID: 35532219 Free PMC article. Review.
-
Improving the pharmacokinetics, biodistribution and plasma stability of monobodies.Front Pharmacol. 2024 Apr 4;15:1393112. doi: 10.3389/fphar.2024.1393112. Front Pharmacol. 2024. PMID: 38617793 Free PMC article.
-
An allosteric inhibitor targeting the STAT3 coiled-coil domain selectively suppresses proliferation of breast cancer.RSC Med Chem. 2025 Mar 24. doi: 10.1039/d4md00926f. Online ahead of print. RSC Med Chem. 2025. PMID: 40190419 Free PMC article.
References
-
- Schindler C, Levy DE, Decker T. JAK-STAT signaling: from interferons to cytokines. J. Biol. Chem. 2007;282:20059–20063. - PubMed
-
- Haura EB, Turkson J, Jove R. Mechanisms of disease: insights into the emerging role of signal transducers and activators of transcription in cancer. Nat. Clin. Pract. Oncol. 2005;2:315–324. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

