A single-cell transcriptome atlas of marsupial embryogenesis and X inactivation
- PMID: 32814901
- PMCID: PMC7116401
- DOI: 10.1038/s41586-020-2629-6
A single-cell transcriptome atlas of marsupial embryogenesis and X inactivation
Erratum in
-
Publisher Correction: A single-cell transcriptome atlas of marsupial embryogenesis and X inactivation.Nature. 2020 Nov;587(7832):E1. doi: 10.1038/s41586-020-2840-5. Nature. 2020. PMID: 33067604
-
Publisher Correction: A single-cell transcriptome atlas of marsupial embryogenesis and X inactivation.Nature. 2021 Feb;590(7844):E28. doi: 10.1038/s41586-020-03110-w. Nature. 2021. PMID: 33469225 No abstract available.
Abstract
Single-cell RNA sequencing of embryos can resolve the transcriptional landscape of development at unprecedented resolution. To date, single-cell RNA-sequencing studies of mammalian embryos have focused exclusively on eutherian species. Analysis of mammalian outgroups has the potential to identify deeply conserved lineage specification and pluripotency factors, and can extend our understanding of X dosage compensation. Metatherian (marsupial) mammals diverged from eutherians around 160 million years ago. They exhibit distinctive developmental features, including late implantation1 and imprinted X chromosome inactivation2, which is associated with expression of the XIST-like noncoding RNA RSX3. Here we perform a single-cell RNA-sequencing analysis of embryogenesis and X chromosome inactivation in a marsupial, the grey short-tailed opossum (Monodelphis domestica). We resolve the developmental trajectory and transcriptional signatures of the epiblast, primitive endoderm and trophectoderm, and identify deeply conserved lineage-specific markers that pre-date the eutherian-marsupial divergence. RSX coating and inactivation of the X chromosome occurs early and rapidly. This observation supports the hypothesis that-in organisms with early X chromosome inactivation-imprinted X chromosome inactivation prevents biallelic X silencing. We identify XSR, an RSX antisense transcript expressed from the active X chromosome, as a candidate for the regulator of imprinted X chromosome inactivation. Our datasets provide insights into the evolution of mammalian embryogenesis and X dosage compensation.
Figures







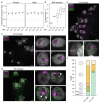
Similar articles
-
Divergent DNA methylation dynamics in marsupial and eutherian embryos.Nature. 2025 Jun;642(8069):1073-1079. doi: 10.1038/s41586-025-08992-2. Epub 2025 May 14. Nature. 2025. PMID: 40369084 Free PMC article.
-
Chromosome-wide profiling of X-chromosome inactivation and epigenetic states in fetal brain and placenta of the opossum, Monodelphis domestica.Genome Res. 2014 Jan;24(1):70-83. doi: 10.1101/gr.161919.113. Epub 2013 Sep 24. Genome Res. 2014. PMID: 24065774 Free PMC article.
-
Rsx is a metatherian RNA with Xist-like properties in X-chromosome inactivation.Nature. 2012 Jul 12;487(7406):254-8. doi: 10.1038/nature11171. Nature. 2012. PMID: 22722828 Free PMC article.
-
Concise review: Pluripotency and the transcriptional inactivation of the female Mammalian X chromosome.Stem Cells. 2012 Jan;30(1):48-54. doi: 10.1002/stem.755. Stem Cells. 2012. PMID: 21997775 Free PMC article. Review.
-
Imprinted X-chromosome inactivation: enlightenment from embryos in vivo.Semin Cell Dev Biol. 2003 Dec;14(6):319-29. doi: 10.1016/j.semcdb.2003.09.027. Semin Cell Dev Biol. 2003. PMID: 15015739 Review.
Cited by
-
Monodelphis domestica Induced Pluripotent Stem Cells Reveal Metatherian Pluripotency Architecture.Int J Mol Sci. 2022 Oct 20;23(20):12623. doi: 10.3390/ijms232012623. Int J Mol Sci. 2022. PMID: 36293487 Free PMC article.
-
Investigation of Food Marking and Contamination Behaviors Informs Feeding Practice in Research Colonies of Laboratory Opossums.J Am Assoc Lab Anim Sci. 2025 Jan 1;64(1):89-97. doi: 10.30802/AALAS-JAALAS-24-060. J Am Assoc Lab Anim Sci. 2025. PMID: 40035216 Free PMC article.
-
Marsupials and Multi-Omics: Establishing New Comparative Models of Neural Crest Patterning and Craniofacial Development.Front Cell Dev Biol. 2022 Jun 23;10:941168. doi: 10.3389/fcell.2022.941168. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 35813210 Free PMC article.
-
A hexa-species transcriptome atlas of mammalian embryogenesis delineates metabolic regulation across three different implantation modes.Nat Commun. 2022 Jun 16;13(1):3407. doi: 10.1038/s41467-022-30194-x. Nat Commun. 2022. PMID: 35710749 Free PMC article.
-
Compensation of gene dosage on the mammalian X.Development. 2024 Aug 1;151(15):dev202891. doi: 10.1242/dev.202891. Epub 2024 Aug 14. Development. 2024. PMID: 39140247 Free PMC article. Review.
References
-
- Sharman GB. Late DNA replication in the paternally derived X chromosome of female kangaroos. Nature. 1971;230:231–232. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

