This is a preprint.
Quantifying absolute neutralization titers against SARS-CoV-2 by a standardized virus neutralization assay allows for cross-cohort comparisons of COVID-19 sera
- PMID: 32817961
- PMCID: PMC7430605
- DOI: 10.1101/2020.08.13.20157222
Quantifying absolute neutralization titers against SARS-CoV-2 by a standardized virus neutralization assay allows for cross-cohort comparisons of COVID-19 sera
Update in
-
Quantifying Absolute Neutralization Titers against SARS-CoV-2 by a Standardized Virus Neutralization Assay Allows for Cross-Cohort Comparisons of COVID-19 Sera.mBio. 2021 Feb 16;12(1):e02492-20. doi: 10.1128/mBio.02492-20. mBio. 2021. PMID: 33593976 Free PMC article.
Abstract
The global COVID-19 pandemic has mobilized efforts to develop vaccines and antibody-based therapeutics, including convalescent plasma therapy, that inhibit viral entry by inducing or transferring neutralizing antibodies (nAbs) against the SARS-CoV-2 spike glycoprotein (CoV2-S). However, rigorous efficacy testing requires extensive screening with live virus under onerous BSL3 conditions which limits high throughput screening of patient and vaccine sera. Myriad BSL-2 compatible surrogate virus neutralization assays (VNAs) have been developed to overcome this barrier. Yet, there is marked variability between VNAs and how their results are presented, making inter-group comparisons difficult. To address these limitations, we developed a standardized VNA using VSVΔG-based CoV-2-S pseudotyped particles (CoV2pp) that can be robustly produced at scale and generate accurate neutralizing titers within 18 hours post-infection. Our standardized CoV2pp VNA showed a strong positive correlation with CoV2-S ELISA and live virus neutralizations in confirmed convalescent patient sera. Three independent groups subsequently validated our standardized CoV2pp VNA (n>120). Our data show that absolute (abs) IC50, IC80, and IC90 values can be legitimately compared across diverse cohorts, highlight the substantial but consistent variability in neutralization potency across these cohorts, and support the use of absIC80 as a more meaningful metric for assessing the neutralization potency of vaccine or convalescent sera. Lastly, we used our CoV2pp in a screen to identify ultra-permissive 293T clones that stably express ACE2 or ACE2+TMPRSS2. When used in combination with our CoV2pp, we can now produce CoV2pp sufficient for 150,000 standardized VNA/week.
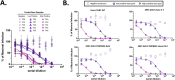
Figures






Similar articles
-
Quantifying Absolute Neutralization Titers against SARS-CoV-2 by a Standardized Virus Neutralization Assay Allows for Cross-Cohort Comparisons of COVID-19 Sera.mBio. 2021 Feb 16;12(1):e02492-20. doi: 10.1128/mBio.02492-20. mBio. 2021. PMID: 33593976 Free PMC article.
-
Pre-Omicron Vaccine Breakthrough Infection Induces Superior Cross-Neutralization against SARS-CoV-2 Omicron BA.1 Compared to Infection Alone.Int J Mol Sci. 2022 Jul 12;23(14):7675. doi: 10.3390/ijms23147675. Int J Mol Sci. 2022. PMID: 35887023 Free PMC article.
-
Comprehensive Comparison of Seven SARS-CoV-2-Specific Surrogate Virus Neutralization and Anti-Spike IgG Antibody Assays Using a Live-Virus Neutralization Assay as a Reference.Microbiol Spectr. 2023 Feb 14;11(1):e0231422. doi: 10.1128/spectrum.02314-22. Epub 2023 Jan 9. Microbiol Spectr. 2023. PMID: 36622205 Free PMC article.
-
SARS-CoV-2 Antibody Neutralization Assay Platforms Based on Epitopes Sources: Live Virus, Pseudovirus, and Recombinant S Glycoprotein RBD.Immune Netw. 2021 Nov 23;21(6):e39. doi: 10.4110/in.2021.21.e39. eCollection 2021 Dec. Immune Netw. 2021. PMID: 35036026 Free PMC article. Review.
-
Advances in Surrogate Neutralization Tests for High-Throughput Screening and the Point-of-Care.Anal Chem. 2025 Mar 18;97(10):5407-5423. doi: 10.1021/acs.analchem.5c00666. Epub 2025 Mar 4. Anal Chem. 2025. PMID: 40035475 Free PMC article. Review. No abstract available.
Cited by
-
Metabolic liver disease - what's in a name?Nat Rev Endocrinol. 2021 Feb;17(2):79-80. doi: 10.1038/s41574-020-00452-3. Nat Rev Endocrinol. 2021. PMID: 33257823 Review. No abstract available.
References
-
- Center for Disease Control. Revised U.S. Surveillance Case Definition for Severe Acute Respiratory Syndrome (SARS) and Update on SARS Cases - United States and Worldwide, December 2003. (2003). - PubMed
-
- World Health Organization. Novel Coronavirus (2019-nCoV): situation report, 19 (2020).
Publication types
Grants and funding
- T32 AI007647/AI/NIAID NIH HHS/United States
- R01 AI123449/AI/NIAID NIH HHS/United States
- HHSN272201400008C/AI/NIAID NIH HHS/United States
- UL1 TR001433/TR/NCATS NIH HHS/United States
- R21 AI148033/AI/NIAID NIH HHS/United States
- I01 BX003860/BX/BLRD VA/United States
- R01 AI116851/AI/NIAID NIH HHS/United States
- IK6 BX004607/BX/BLRD VA/United States
- F31 HL149295/HL/NHLBI NIH HHS/United States
- R21 AI149033/AI/NIAID NIH HHS/United States
- R01 AI143839/AI/NIAID NIH HHS/United States
- F31 AI154739/AI/NIAID NIH HHS/United States
- 75N93019C00051/AI/NIAID NIH HHS/United States
- R01 AI139290/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
