Assembly of a Ribozyme Ligase from Short Oligomers by Nonenzymatic Ligation
- PMID: 32820909
- PMCID: PMC9594310
- DOI: 10.1021/jacs.0c06722
Assembly of a Ribozyme Ligase from Short Oligomers by Nonenzymatic Ligation
Abstract
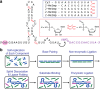
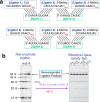
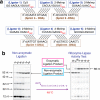
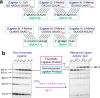
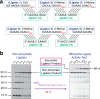
Our current understanding of the chemistry of the primordial genetic material is fragmentary at best. The chemical replication of oligonucleotides long enough to perform catalytic functions is particularly problematic because of the low efficiency of nonenzymatic template copying. Here we show that this problem can be circumvented by assembling a functional ribozyme by the templated ligation of short oligonucleotides. However, this approach creates a new problem because the splint oligonucleotides used to drive ribozyme assembly strongly inhibit the resulting ribozyme. We explored three approaches to the design of splint oligonucleotides that enable efficient ligation but which allow the assembled ribozyme to remain active. DNA splints, splints with G:U wobble pairs, and splints with G to I (Inosine) substitutions all allowed for the efficient assembly of an active ribozyme ligase. Our work demonstrates the possibility of a transition from nonenzymatic ligation to enzymatic ligation and reveals the importance of avoiding ribozyme inhibition by complementary oligonucleotides.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
Similar articles
-
In vitro adaptation of a ligase ribozyme for activity under a low-pH condition.Biotechnol Bioeng. 2001 Dec 5;75(5):590-6. doi: 10.1002/bit.10033. Biotechnol Bioeng. 2001. PMID: 11745135
-
Isolation of a ribozyme with 5'-5' ligase activity.Chem Biol. 1995 May;2(5):325-33. doi: 10.1016/1074-5521(95)90051-9. Chem Biol. 1995. PMID: 9383434
-
Nonenzymatic copying of RNA templates containing all four letters is catalyzed by activated oligonucleotides.Elife. 2016 Jun 28;5:e17756. doi: 10.7554/eLife.17756. Elife. 2016. PMID: 27351102 Free PMC article.
-
Regulation of ribozyme activity with short oligonucleotides.Biol Pharm Bull. 2004 Apr;27(4):457-62. doi: 10.1248/bpb.27.457. Biol Pharm Bull. 2004. PMID: 15056847 Review.
-
Structure and function of the hairpin ribozyme.J Mol Biol. 2000 Mar 24;297(2):269-91. doi: 10.1006/jmbi.2000.3560. J Mol Biol. 2000. PMID: 10715200 Review.
Cited by
-
Origin of the RNA World in Cold Hadean Geothermal Fields Enriched in Zinc and Potassium: Abiogenesis as a Positive Fallout from the Moon-Forming Impact?Life (Basel). 2025 Mar 4;15(3):399. doi: 10.3390/life15030399. Life (Basel). 2025. PMID: 40141744 Free PMC article.
-
Nonenzymatic assembly of active chimeric ribozymes from aminoacylated RNA oligonucleotides.Proc Natl Acad Sci U S A. 2022 Feb 15;119(7):e2116840119. doi: 10.1073/pnas.2116840119. Proc Natl Acad Sci U S A. 2022. PMID: 35140183 Free PMC article.
-
Experimental Tests of the Virtual Circular Genome Model for Nonenzymatic RNA Replication.J Am Chem Soc. 2023 Apr 5;145(13):7504-7515. doi: 10.1021/jacs.3c00612. Epub 2023 Mar 24. J Am Chem Soc. 2023. PMID: 36963403 Free PMC article.
-
On the origin of life: an RNA-focused synthesis and narrative.RNA. 2023 Aug;29(8):1085-1098. doi: 10.1261/rna.079598.123. Epub 2023 May 4. RNA. 2023. PMID: 37142437 Free PMC article.
-
Enhanced nonenzymatic RNA copying with in-situ activation of short oligonucleotides.Nucleic Acids Res. 2023 Jul 21;51(13):6528-6539. doi: 10.1093/nar/gkad439. Nucleic Acids Res. 2023. PMID: 37247941 Free PMC article.
References
-
- Woese C. R.The Genetic Code the Molecular Basis for Genetic Expression; Harper & Row: New York, 1967.
-
- Gilbert W. Origin of Life: The RNA World. Nature 1986, 319 (6055), 618–618. 10.1038/319618a0. - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

