COVID-19 pandemic: Insights into structure, function, and hACE2 receptor recognition by SARS-CoV-2
- PMID: 32822426
- PMCID: PMC7444525
- DOI: 10.1371/journal.ppat.1008762
COVID-19 pandemic: Insights into structure, function, and hACE2 receptor recognition by SARS-CoV-2
Abstract
Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) is a newly emerging, highly transmissible, and pathogenic coronavirus in humans that has caused global public health emergencies and economic crises. To date, millions of infections and thousands of deaths have been reported worldwide, and the numbers continue to rise. Currently, there is no specific drug or vaccine against this deadly virus; therefore, there is a pressing need to understand the mechanism(s) through which this virus enters the host cell. Viral entry into the host cell is a multistep process in which SARS-CoV-2 utilizes the receptor-binding domain (RBD) of the spike (S) glycoprotein to recognize angiotensin-converting enzyme 2 (ACE2) receptors on the human cells; this initiates host-cell entry by promoting viral-host cell membrane fusion through large-scale conformational changes in the S protein. Receptor recognition and fusion are critical and essential steps of viral infections and are key determinants of the viral host range and cross-species transmission. In this review, we summarize the current knowledge on the origin and evolution of SARS-CoV-2 and the roles of key viral factors. We discuss the structure of RNA-dependent RNA polymerase (RdRp) of SARS-CoV-2 and its significance in drug discovery and explain the receptor recognition mechanisms of coronaviruses. Further, we provide a comparative analysis of the SARS-CoV and SARS-CoV-2 S proteins and their receptor-binding specificity and discuss the differences in their antigenicity based on biophysical and structural characteristics.
Conflict of interest statement
"The authors have declared that no competing interests exist".
Figures


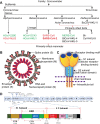


References
-
- WHO. Coronavirus never before seen in humans is the cause of SARS [Internet]. 2003 [cited 2020 Jun 27]. Available from: https://www.who.int/mediacentre/news/releases/2003/pr31/en/
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

