Efficient genome editing in pathogenic mycobacteria using Streptococcus thermophilus CRISPR1-Cas9
- PMID: 32829077
- PMCID: PMC7612230
- DOI: 10.1016/j.tube.2020.101983
Efficient genome editing in pathogenic mycobacteria using Streptococcus thermophilus CRISPR1-Cas9
Abstract
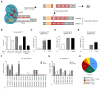
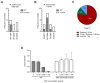
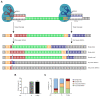
The ability to genetically engineer pathogenic mycobacteria has increased significantly over the last decades due to the generation of new molecular tools. Recently, the application of the Streptococcus pyogenes and the Streptococcus thermophilus CRISPR-Cas9 systems in mycobacteria has enabled gene editing and efficient CRISPR interference-mediated transcriptional regulation. Here, we converted CRISPR interference into an efficient genome editing tool for mycobacteria. We demonstrate that the Streptococcus thermophilus CRISPR1-Cas9 (Sth1Cas9) is functional in Mycobacterium marinum and Mycobacterium tuberculosis, enabling highly efficient and precise DNA breaks and indel formation, without any off-target effects. In addition, with dual sgRNAs this system can be used to generate two indels simultaneously or to create specific deletions. The ability to use the power of the CRISPR-Cas9-mediated gene editing toolbox in M. tuberculosis with a single step will accelerate research into this deadly pathogen.
Keywords: CRISPR-Cas9 system; Genome editing; Indels; Mycobacterium marinum; Mycobacterium tuberculosis.
Copyright © 2020 The Author(s). Published by Elsevier Ltd.. All rights reserved.
Conflict of interest statement
None
Figures
Similar articles
-
A CRISPR-Assisted Nonhomologous End-Joining Strategy for Efficient Genome Editing in Mycobacterium tuberculosis.mBio. 2020 Jan 28;11(1):e02364-19. doi: 10.1128/mBio.02364-19. mBio. 2020. PMID: 31992616 Free PMC article.
-
Versatile and robust genome editing with Streptococcus thermophilus CRISPR1-Cas9.Genome Res. 2020 Jan;30(1):107-117. doi: 10.1101/gr.255414.119. Epub 2020 Jan 3. Genome Res. 2020. PMID: 31900288 Free PMC article.
-
Programmable transcriptional repression in mycobacteria using an orthogonal CRISPR interference platform.Nat Microbiol. 2017 Feb 6;2:16274. doi: 10.1038/nmicrobiol.2016.274. Nat Microbiol. 2017. PMID: 28165460 Free PMC article.
-
CRISPR/Cas9 gene-editing strategies in cardiovascular cells.Cardiovasc Res. 2020 Apr 1;116(5):894-907. doi: 10.1093/cvr/cvz250. Cardiovasc Res. 2020. PMID: 31584620 Review.
-
Optimization of genome editing through CRISPR-Cas9 engineering.Bioengineered. 2016 Apr;7(3):166-74. doi: 10.1080/21655979.2016.1189039. Bioengineered. 2016. PMID: 27340770 Free PMC article. Review.
Cited by
-
CRISPY-BRED and CRISPY-BRIP: efficient bacteriophage engineering.Sci Rep. 2021 Mar 24;11(1):6796. doi: 10.1038/s41598-021-86112-6. Sci Rep. 2021. PMID: 33762639 Free PMC article.
-
A dual-plasmid CRISPR/Cas9-based method for rapid and efficient genetic disruption in Mycobacterium abscessus.J Bacteriol. 2024 Mar 21;206(3):e0033523. doi: 10.1128/jb.00335-23. Epub 2024 Feb 6. J Bacteriol. 2024. PMID: 38319218 Free PMC article.
-
Phage engineering and phage-assisted CRISPR-Cas delivery to combat multidrug-resistant pathogens.Bioeng Transl Med. 2022 Aug 6;8(2):e10381. doi: 10.1002/btm2.10381. eCollection 2023 Mar. Bioeng Transl Med. 2022. PMID: 36925687 Free PMC article. Review.
-
An intramolecular cross-talk in D29 mycobacteriophage endolysin governs the lytic cycle and phage-host population dynamics.Sci Adv. 2024 Feb 9;10(6):eadh9812. doi: 10.1126/sciadv.adh9812. Epub 2024 Feb 9. Sci Adv. 2024. PMID: 38335296 Free PMC article.
-
The mevalonate pathway of isoprenoid biosynthesis supports metabolic flexibility in Mycobacterium marinum.bioRxiv [Preprint]. 2025 Jul 11:2025.07.11.664281. doi: 10.1101/2025.07.11.664281. bioRxiv. 2025. PMID: 40672248 Free PMC article. Preprint.
References
-
- WHO. Global Tuberculosis Report. 2019
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

