Fruitful Neutralizing Antibody Pipeline Brings Hope To Defeat SARS-Cov-2
- PMID: 32829936
- PMCID: PMC7572790
- DOI: 10.1016/j.tips.2020.07.004
Fruitful Neutralizing Antibody Pipeline Brings Hope To Defeat SARS-Cov-2
Abstract
With the recent spread of severe acute respiratory syndrome coronavirus (SARS-CoV-2)_ infecting >16 million people worldwide as of 28 July 2020, causing >650 000 deaths, there is a desperate need for therapeutic agents and vaccines. Building on knowledge of previous outbreaks of SARS-CoV-1 and Middle East respiratory syndrome (MERS), the development of therapeutic antibodies and vaccines against coronavirus disease 2019 (COVID-19) is taking place at an unprecedented speed. Current efforts towards the development of neutralizing antibodies against COVID-19 are summarized. We also highlight the importance of a fruitful antibody development pipeline to combat the potential escape plans of SARS-CoV-2, including somatic mutations and antibody-dependent enhancement (ADE).
Keywords: COVID-19; SARS-CoV-2; antibody-dependent enhancement; drug resistance; neutralizing antibodies; viral escape.
Published by Elsevier Ltd.
Figures



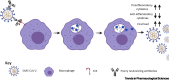
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

