Dynamic incorporation of multiple in silico functional annotations empowers rare variant association analysis of large whole-genome sequencing studies at scale
- PMID: 32839606
- PMCID: PMC7483769
- DOI: 10.1038/s41588-020-0676-4
Dynamic incorporation of multiple in silico functional annotations empowers rare variant association analysis of large whole-genome sequencing studies at scale
Abstract
Large-scale whole-genome sequencing studies have enabled the analysis of rare variants (RVs) associated with complex phenotypes. Commonly used RV association tests have limited scope to leverage variant functions. We propose STAAR (variant-set test for association using annotation information), a scalable and powerful RV association test method that effectively incorporates both variant categories and multiple complementary annotations using a dynamic weighting scheme. For the latter, we introduce 'annotation principal components', multidimensional summaries of in silico variant annotations. STAAR accounts for population structure and relatedness and is scalable for analyzing very large cohort and biobank whole-genome sequencing studies of continuous and dichotomous traits. We applied STAAR to identify RVs associated with four lipid traits in 12,316 discovery and 17,822 replication samples from the Trans-Omics for Precision Medicine Program. We discovered and replicated new RV associations, including disruptive missense RVs of NPC1L1 and an intergenic region near APOC1P1 associated with low-density lipoprotein cholesterol.
Figures
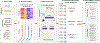

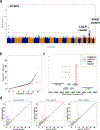
References
-
- Morgenthaler S & Thilly WG A strategy to discover genes that carry multi-allelic or mono-allelic risk for common diseases: a cohort allelic sums test (CAST). Mutation Research/Fundamental and Molecular Mechanisms of Mutagenesis 615, 28–56 (2007). - PubMed
Methods-only references
Publication types
MeSH terms
Substances
Grants and funding
- U19 CA203654/CA/NCI NIH HHS/United States
- R35 HL135824/HL/NHLBI NIH HHS/United States
- N01 HC095160/HL/NHLBI NIH HHS/United States
- U01 HL054472/HL/NHLBI NIH HHS/United States
- R01 HL071025/HL/NHLBI NIH HHS/United States
- R01 HL112064/HL/NHLBI NIH HHS/United States
- 75N92020D00002/HL/NHLBI NIH HHS/United States
- HHSN268201500003C/HL/NHLBI NIH HHS/United States
- R01 HL113323/HL/NHLBI NIH HHS/United States
- N01 HC095161/HL/NHLBI NIH HHS/United States
- P30 CA016672/CA/NCI NIH HHS/United States
- 75N92020D00005/HL/NHLBI NIH HHS/United States
- R01 HL104135/HL/NHLBI NIH HHS/United States
- R35 CA197449/CA/NCI NIH HHS/United States
- P30 ES010126/ES/NIEHS NIH HHS/United States
- HHSN268201800012C/HL/NHLBI NIH HHS/United States
- N01 HC095168/HL/NHLBI NIH HHS/United States
- R01 HL133040/HL/NHLBI NIH HHS/United States
- R01 HL120393/HL/NHLBI NIH HHS/United States
- R01 HL087698/HL/NHLBI NIH HHS/United States
- R03 HL141439/HL/NHLBI NIH HHS/United States
- K01 AG059898/AG/NIA NIH HHS/United States
- U01 DK085524/DK/NIDDK NIH HHS/United States
- HHSN268201600002C/HL/NHLBI NIH HHS/United States
- HHSN268201500001C/HL/NHLBI NIH HHS/United States
- UL1 TR001079/TR/NCATS NIH HHS/United States
- T32 GM135117/GM/NIGMS NIH HHS/United States
- N01 HC095169/HL/NHLBI NIH HHS/United States
- HHSN268201600018C/HL/NHLBI NIH HHS/United States
- HHSN268201800014I/HB/NHLBI NIH HHS/United States
- U01 HL130114/HL/NHLBI NIH HHS/United States
- R01 AR048797/AR/NIAMS NIH HHS/United States
- R01 HL092577/HL/NHLBI NIH HHS/United States
- P30 ES000002/ES/NIEHS NIH HHS/United States
- P50 HL118006/HL/NHLBI NIH HHS/United States
- U01 HL054509/HL/NHLBI NIH HHS/United States
- 75N92020D00001/HL/NHLBI NIH HHS/United States
- R01 HL113338/HL/NHLBI NIH HHS/United States
- R01 AG058921/AG/NIA NIH HHS/United States
- R01 NS058700/NS/NINDS NIH HHS/United States
- T32 HG000040/HG/NHGRI NIH HHS/United States
- N01 HC095167/HL/NHLBI NIH HHS/United States
- U01 HL137181/HL/NHLBI NIH HHS/United States
- HHSN268201800014C/HL/NHLBI NIH HHS/United States
- N01 HC095159/HL/NHLBI NIH HHS/United States
- 75N92020D00003/HL/NHLBI NIH HHS/United States
- F32 HL085989/HL/NHLBI NIH HHS/United States
- R01 MH078111/MH/NIMH NIH HHS/United States
- R01 HL119443/HL/NHLBI NIH HHS/United States
- R01 HL105756/HL/NHLBI NIH HHS/United States
- K01 HL125751/HL/NHLBI NIH HHS/United States
- R01 HL067348/HL/NHLBI NIH HHS/United States
- R01 HL142711/HL/NHLBI NIH HHS/United States
- R35 HL135818/HL/NHLBI NIH HHS/United States
- T32 GM074897/GM/NIGMS NIH HHS/United States
- P30 DK063491/DK/NIDDK NIH HHS/United States
- U01 HL072524/HL/NHLBI NIH HHS/United States
- HHSN268201700002C/HL/NHLBI NIH HHS/United States
- K08 HL141601/HL/NHLBI NIH HHS/United States
- HHSN268201800001C/HL/NHLBI NIH HHS/United States
- HHSN268201700001I/HL/NHLBI NIH HHS/United States
- N01 HC025195/HL/NHLBI NIH HHS/United States
- HHSN268201800013I/MD/NIMHD NIH HHS/United States
- HHSN268201600003C/HL/NHLBI NIH HHS/United States
- HHSN268201700004I/HL/NHLBI NIH HHS/United States
- R35 GM127131/GM/NIGMS NIH HHS/United States
- HHSN268201500001I/HL/NHLBI NIH HHS/United States
- HHSN268201800012I/HL/NHLBI NIH HHS/United States
- M01 RR000052/RR/NCRR NIH HHS/United States
- UL1 TR001420/TR/NCATS NIH HHS/United States
- R01 HL049762/HL/NHLBI NIH HHS/United States
- 75N92020D00004/HL/NHLBI NIH HHS/United States
- HHSN268201600004C/HL/NHLBI NIH HHS/United States
- P01 HL045522/HL/NHLBI NIH HHS/United States
- 75N92020D00007/HL/NHLBI NIH HHS/United States
- U01 HL072518/HL/NHLBI NIH HHS/United States
- N01 HC095163/HL/NHLBI NIH HHS/United States
- HHSN268201800011C/HL/NHLBI NIH HHS/United States
- U01 HL137162/HL/NHLBI NIH HHS/United States
- M01 RR007122/RR/NCRR NIH HHS/United States
- HHSN268201500003I/HL/NHLBI NIH HHS/United States
- HHSN268201600001C/HL/NHLBI NIH HHS/United States
- R01 HL059684/HL/NHLBI NIH HHS/United States
- R01 HL093093/HL/NHLBI NIH HHS/United States
- HHSN268201700005C/HL/NHLBI NIH HHS/United States
- R01 DK071891/DK/NIDDK NIH HHS/United States
- U24 HG009446/HG/NHGRI NIH HHS/United States
- HHSN268201700003C/HL/NHLBI NIH HHS/United States
- HHSN268201700001C/HL/NHLBI NIH HHS/United States
- R01 MH078143/MH/NIMH NIH HHS/United States
- U24 CA237617/CA/NCI NIH HHS/United States
- HHSN268201800015I/HB/NHLBI NIH HHS/United States
- 75N92019D00031/HL/NHLBI NIH HHS/United States
- HHSN268201700004C/HL/NHLBI NIH HHS/United States
- UL1 TR000040/TR/NCATS NIH HHS/United States
- P01 CA134294/CA/NCI NIH HHS/United States
- N01 HC095166/HL/NHLBI NIH HHS/United States
- HHSN268201700002I/HL/NHLBI NIH HHS/United States
- R01 MH101244/MH/NIMH NIH HHS/United States
- R01 MH083824/MH/NIMH NIH HHS/United States
- HHSN268201800010I/HB/NHLBI NIH HHS/United States
- HHSN268201700005I/HL/NHLBI NIH HHS/United States
- U01 HG009088/HG/NHGRI NIH HHS/United States
- P42 ES016454/ES/NIEHS NIH HHS/United States
- 75N92020D00006/HL/NHLBI NIH HHS/United States
- R01 HL117626/HL/NHLBI NIH HHS/United States
- N01 HC095162/HL/NHLBI NIH HHS/United States
- UL1 TR001881/TR/NCATS NIH HHS/United States
- HHSN268201800011I/HB/NHLBI NIH HHS/United States
- R01 HL134320/HL/NHLBI NIH HHS/United States
- N01 HC095164/HL/NHLBI NIH HHS/United States
- N01 HC095165/HL/NHLBI NIH HHS/United States
- R13 CA124365/CA/NCI NIH HHS/United States
- HHSN268201700003I/HL/NHLBI NIH HHS/United States
- U01 HL054495/HL/NHLBI NIH HHS/United States
- R01 HL055673/HL/NHLBI NIH HHS/United States
- R01 HL092301/HL/NHLBI NIH HHS/United States
- U01 HL054473/HL/NHLBI NIH HHS/United States

