Upregulation of METTL14 mediates the elevation of PERP mRNA N6 adenosine methylation promoting the growth and metastasis of pancreatic cancer
- PMID: 32843065
- PMCID: PMC7446161
- DOI: 10.1186/s12943-020-01249-8
Upregulation of METTL14 mediates the elevation of PERP mRNA N6 adenosine methylation promoting the growth and metastasis of pancreatic cancer
Abstract
Background: Pancreatic cancer is one of the most lethal human cancers. N6-methyladenosine (m6A), a common eukaryotic mRNA modification, plays critical roles in both physiological and pathological processes. However, its role in pancreatic cancer remains elusive.
Methods: LC/MS was used to profile m6A levels in pancreatic cancer and normal tissues. Bioinformatics analysis, real-time PCR, immunohistochemistry, and western blotting were used to identify the role of m6A regulators in pancreatic cancer. The biological effects of methyltransferase-like 14 (METTL14), an mRNA methylase, were investigated using in vitro and in vivo models. MeRIP-Seq and RNA-Seq were used to assess the downstream targets of METTL14.
Results: We found that the m6A levels were elevated in approximately 70% of the pancreatic cancer samples. Furthermore, we demonstrated that METTL14 is the major enzyme that modulates m6A methylation (frequency and site of methylation). METTL14 overexpression markedly promoted pancreatic cancer cell proliferation and migration both in vitro and in vivo, via direct targeting of the downstream PERP mRNA (p53 effector related to PMP-22) in an m6A-dependent manner. Methylation of the target adenosine lead to increased PERP mRNA turnover, thus decreasing PERP (mRNA and protein) levels in pancreatic cancer cells.
Conclusions: Our data suggest that the upregulation of METTL14 leads to the decrease of PERP levels via m6A modification, promoting the growth and metastasis of pancreatic cancer; therefore METTL14 is a potential therapeutic target for its treatment.
Keywords: METTL14; N6-methyladenosine; PERP; Pancreatic cancer; m6A.
Conflict of interest statement
C.H. is the scientific founder of Accent Therapeutics and a member of its scientific advisory board. All other authors declare no competing financial interests.
Figures



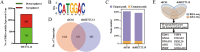


References
-
- Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108. - PubMed
-
- Middleton G, Palmer DH, Greenhalf W, Ghaneh P, Jackson R, Cox T, et al. Vandetanib plus gemcitabine versus placebo plus gemcitabine in locally advanced or metastatic pancreatic carcinoma (ViP): a prospective, randomised, double-blind, multicentre phase 2 trial. Lancet Oncol. 2017;18:486–499. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

