XK is a partner for VPS13A: a molecular link between Chorea-Acanthocytosis and McLeod Syndrome
- PMID: 32845802
- PMCID: PMC7851852
- DOI: 10.1091/mbc.E19-08-0439-T
XK is a partner for VPS13A: a molecular link between Chorea-Acanthocytosis and McLeod Syndrome
Abstract
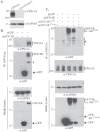
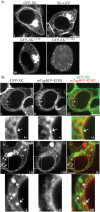
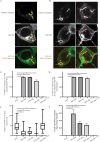
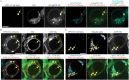
Vps13 is a highly conserved lipid transfer protein found at multiple interorganelle membrane contact sites where it mediates distinct processes. In yeast, recruitment of Vps13 to different contact sites occurs via various partner proteins. In humans, four VPS13 family members, A-D, are associated with different diseases. In particular, vps13A mutants result in the neurodegenerative disorder Chorea-Acanthocytosis (ChAc). ChAc phenotypes resemble those of McLeod Syndrome, caused by mutations in the XK gene, suggesting that XK could be a partner protein for VPS13A. XK does, in fact, exhibit hallmarks of a VPS13A partner: it forms a complex with VPS13A in human cells and, when overexpressed, relocalizes VPS13A from lipid droplets to subdomains of the endoplasmic reticulum. Introduction of two different ChAc disease-linked missense mutations into VPS13A prevents this XK-induced relocalization. These results suggest that dysregulation of a VPS13A-XK complex is the common basis for ChAc and McLeod Syndrome.
Figures



References
-
- Dobson-Stone C, Danek A, Rampoldi L, Hardie RJ, Chalmers RM, Wood NW, Bohlega S, Dotti MT, Federico A, Shizuka M, et al (2002). Mutational spectrum of the CHAC gene in patients with chorea-acanthocytosis. Eur J Hum Genet , 773–781. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

